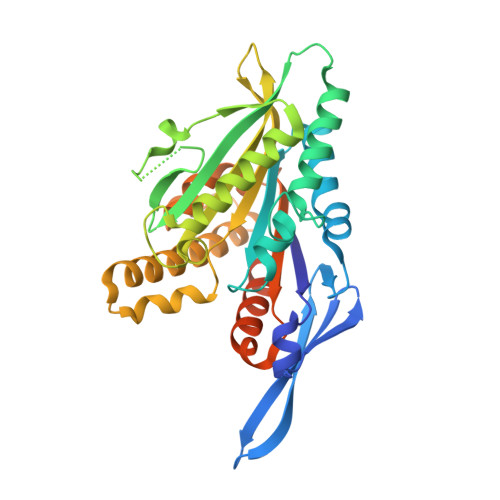
Structure of a kinesin microtubule depolymerization machine.
Shipley, K., Hekmat-Nejad, M., Turner, J., Moores, C., Anderson, R., Milligan, R., Sakowicz, R., Fletterick, R.(2004) EMBO J 23: 1422-1432
- PubMed: 15029249
- DOI: https://doi.org/10.1038/sj.emboj.7600165
- Primary Citation of Related Structures:
1RY6 - PubMed Abstract:
With their ability to depolymerize microtubules (MTs), KinI kinesins are the rogue members of the kinesin family. Here we present the 1.6 A crystal structure of a KinI motor core from Plasmodium falciparum, which is sufficient for depolymerization in vitro. Unlike all published kinesin structures to date, nucleotide is not present, and there are noticeable differences in loop regions L6 and L10 (the plus-end tip), L2 and L8 and in switch II (L11 and helix4); otherwise, the pKinI structure is very similar to previous kinesin structures. KinI-conserved amino acids were mutated to alanine, and studied for their effects on depolymerization and ATP hydrolysis. Notably, mutation of three residues in L2 appears to primarily affect depolymerization, rather than general MT binding or ATP hydrolysis. The results of this study confirm the suspected importance of loop 2 for KinI function, and provide evidence that KinI is specialized to hydrolyze ATP after initiating depolymerization.
Organizational Affiliation:
Graduate Group in Biophysics, University of California, San Francisco, CA, USA.