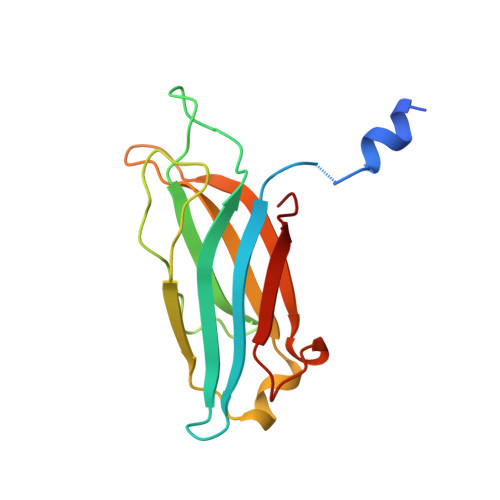
Structure of the first C2 domain of synaptotagmin I: a novel Ca2+/phospholipid-binding fold.
Sutton, R.B., Davletov, B.A., Berghuis, A.M., Sudhof, T.C., Sprang, S.R.(1995) Cell 80: 929-938
- PubMed: 7697723
- DOI: https://doi.org/10.1016/0092-8674(95)90296-1
- Primary Citation of Related Structures:
1RSY - PubMed Abstract:
C2 domains are regulatory sequence motifs that occur widely in nature. Synaptotagmin I, a synaptic vesicle protein involved in the Ca2+ regulation of exocytosis, contains two C2 domains, the first of which acts as a Ca2+ sensor. We now describe the three-dimensional structure of this C2 domain at 1.9 A resolution in both the Ca(2+)-bound and Ca(2+)-free forms. The C2 polypeptide forms an eight-stranded beta sandwich constructed around a conserved four-stranded motif designated as a C2 key. Ca2+ binds in a cup-shaped depression between two polypeptide loops located at the N- and C-termini of the C2-key motif.
Organizational Affiliation:
Department of Biochemistry, University of Texas Southwestern Medical Center, Dallas 75235.