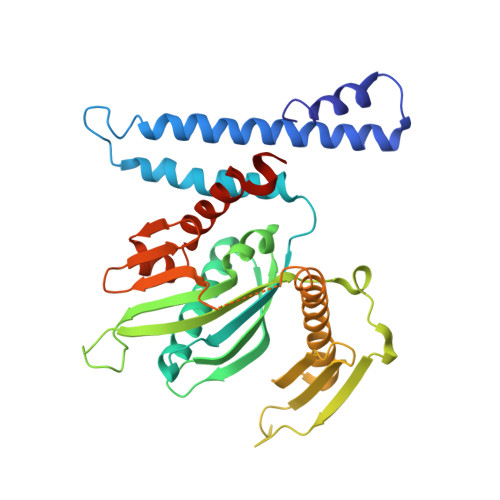
Structural analyses of peptide release factor 1 from Thermotoga maritima reveal domain flexibility required for its interaction with the ribosome.
Shin, D.H., Brandsen, J., Jancarik, J., Yokota, H., Kim, R., Kim, S.H.(2004) J Mol Biol 341: 227-239
- PubMed: 15312775
- DOI: https://doi.org/10.1016/j.jmb.2004.05.055
- Primary Citation of Related Structures:
1RQ0 - PubMed Abstract:
We have determined the crystal structure of peptide chain release factor 1 (RF1) from Thermotoga maritima (gi 4981173) at 2.65 Angstrom resolution by selenomethionine single-wavelength anomalous dispersion (SAD) techniques. RF1 is a protein that recognizes stop codons and promotes the release of a nascent polypeptide from tRNA on the ribosome. Selenomethionine-labeled RF1 crystallized in space group P2(1) with three monomers per asymmetric unit. It has approximate dimensions of 75 Angstrom x 70 Angstrom x 45 Angstrom and is composed of four domains. The overall fold of each RF1 domain shows almost the same topology with Escherichia coli RF2, except that the RF1 N-terminal domain is shorter and the C-terminal domain is longer than that of RF2. The N-terminal domain of RF1 indicates a rigid-body movement relative to that of RF2 with an angle of approximately 90 degrees. Including these features, RF1 has a tripeptide anticodon PVT motif instead of the SPF motif of RF2, which confers the specificity towards the stop codons. The analyses of three molecules in the asymmetric unit and comparison with RF2 revealed the presence of dynamic movement of domains I and III, which are anchored to the central domain by hinge loops. The crystal structure of RF1 elucidates the intrinsic property of this family of having large domain movements for proper function with the ribosome.
Organizational Affiliation:
Physical Biosciences Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA.