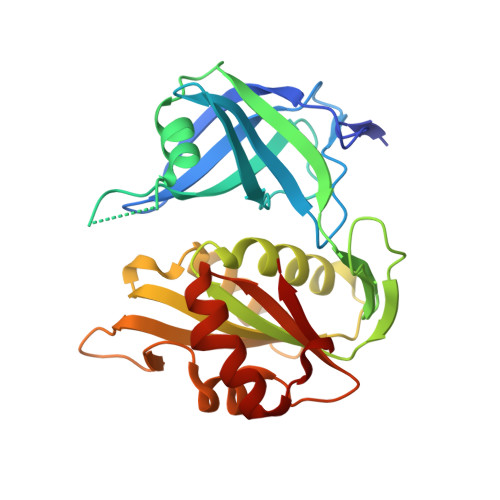
The structure of the S127P mutant of cytochrome b5 reductase that causes methemoglobinemia shows the AMP moiety of the flavin occupying the substrate binding site
Bewley, M.C., Davis, C.A., Marohnic, C.C., Taormina, D., Barber, M.J.(2003) Biochemistry 42: 13145-13151
- PubMed: 14609324
- DOI: https://doi.org/10.1021/bi034915c
- Primary Citation of Related Structures:
1QX4 - PubMed Abstract:
Methemoglobinemia, the first hereditary disease to be identified that involved an enzyme deficiency, has been ascribed to mutations in the enzyme cytochrome b(5) reductase. A variety of defects in either the erythrocytic or microsomal forms of the enzyme have been identified that give rise to the type I or type II variant of the disease, respectively. The positions of the methemoglobinemia-causing mutations are scattered throughout the protein sequence, but the majority of the nontruncated mutants that produce type II symptoms occur close to the flavin adenine dinucleotide (FAD) cofactor binding site. While X-ray structures have been determined for the soluble, flavin-containing diaphorase domains of the rat and pig enzymes, no X-ray or NMR structure has been described for the human enzyme or any of the methemoglobinemia variants. S127P, a mutant that causes type II methemoglobinemia, was the first to be positively identified and have its spectroscopic and kinetic properties characterized that revealed altered nicotinamide adenine dinucleotide hydride (NADH) substrate binding behavior. To understand these changes at a structural level, we have determined the structure of the S127P mutant of rat cytochrome b(5) reductase to 1.8 A resolution, providing the first structural snapshot of a cytochrome b(5) reductase mutant that causes methemoglobinemia. The high-resolution structure revealed that the adenosine diphosphate (ADP) moiety of the FAD prosthetic group is displaced into the corresponding ADP binding site of the physiological substrate, NADH, thus acting as a substrate inhibitor which is consistent with both the spectroscopic and kinetic data.
Organizational Affiliation:
Biology Department, Brookhaven National Laboratory, Upton, New York 11973, USA. mcb21@psu.edu