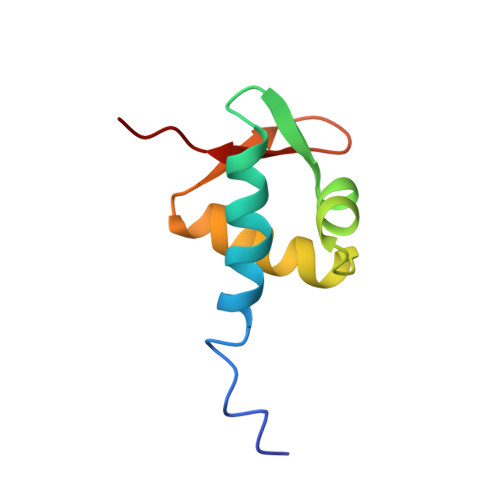
The solution structure of the Zalpha domain of the human RNA editing enzyme ADAR1 reveals a prepositioned binding surface for Z-DNA.
Schade, M., Turner, C.J., Kuhne, R., Schmieder, P., Lowenhaupt, K., Herbert, A., Rich, A., Oschkinat, H.(1999) Proc Natl Acad Sci U S A 96: 12465-12470
- PubMed: 10535945
- DOI: https://doi.org/10.1073/pnas.96.22.12465
- Primary Citation of Related Structures:
1QGP - PubMed Abstract:
Double-stranded RNA deaminase I (ADAR1) contains the Z-DNA binding domain Zalpha. Here we report the solution structure of free Zalpha and map the interaction surface with Z-DNA, confirming roles previously assigned to residues by mutagenesis. Comparison with the crystal structure of the (Zalpha)(2)/Z-DNA complex shows that most Z-DNA contacting residues in free Zalpha are prepositioned to bind Z-DNA, thus minimizing the entropic cost of binding. Comparison with homologous (alpha+beta)helix-turn-helix/B-DNA complexes suggests that binding of Zalpha to B-DNA is disfavored by steric hindrance, but does not eliminate the possibility that related domains may bind to both B- and Z-DNA.
Organizational Affiliation:
Department of Biology, Massachusetts Institute of Technology, Cambridge, MA 02139, USA.