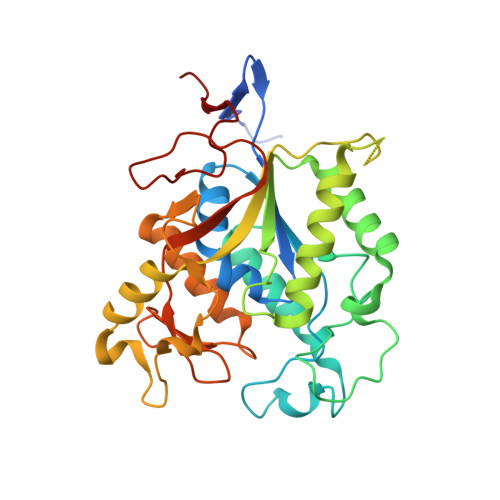
Crystal Structure of MshB from Mycobacterium tuberculosis, a Deacetylase Involved in Mycothiol Biosynthesis.
McCarthy, A.A., Peterson, N.A., Knijff, R., Baker, E.N.(2004) J Mol Biol 335: 1131-1141
- PubMed: 14698305
- DOI: https://doi.org/10.1016/j.jmb.2003.11.034
- Primary Citation of Related Structures:
1Q7T - PubMed Abstract:
All living species require protection against the damaging effects of the reactive oxygen species that are a natural by-product of aerobic life. In most organisms, glutathione is a critical component of these defences, maintaining a reducing environment inside cells. Some bacteria, however, including pathogenic mycobacteria, use an alternative low molecular mass thiol compound called mycothiol (MSH) for this purpose. Enzymes that synthesize MSH are attractive candidates for the design of novel anti-TB drugs because of the importance of MSH for mycobacterial life and the absence of such enzymes in humans. We have determined the three-dimensional structure of MshB (Rv1170), a metal-dependent deacetylase from Mycobacterium tuberculosis that catalyses the second step in MSH biosynthesis. The structure, determined at 1.9A resolution by X-ray crystallography (R=19.0%, R(free)=21.4%), reveals an alpha/beta fold in which helices pack against a seven-stranded mostly parallel beta-sheet. Large loops emanating from the C termini of the beta-strands enclose a deep cavity, which is the location of the putative active site. At the bottom of this cavity is a metal-binding site associated with a sequence motif AHPDDE that is invariant in all homologues. An adventitiously bound beta-octylglucoside molecule, used in crystallization, enables us to model the binding of the true substrate and propose a metal-dependent mechanistic model for deacetylation. Sequence comparisons indicate that MshB is representative of a wider family of enzymes that act on substituted N-acetylglucosamine residues, including a deacetylase involved in the biosynthesis of glycosylphosphatidylinositol (GPI) anchors in eukaryotes.
Organizational Affiliation:
School of Biological Sciences, University of Auckland, Private Bag 92-019, 1, Auckland, New Zealand