The Crystal Structure of 1-D-myo-Inositol 2-Acetamido-2-deoxy-alpha-D-glucopyranoside Deacetylase (MshB) from Mycobacterium tuberculosis Reveals a Zinc Hydrolase with a Lactate Dehydrogenase Fold.
Maynes, J.T., Garen, C., Cherney, M.M., Newton, G., Arad, D., Av-Gay, Y., Fahey, R.C., James, M.N.(2003) J Biol Chem 278: 47166-47170
- PubMed: 12958317
- DOI: https://doi.org/10.1074/jbc.M308914200
- Primary Citation of Related Structures:
1Q74 - PubMed Abstract:
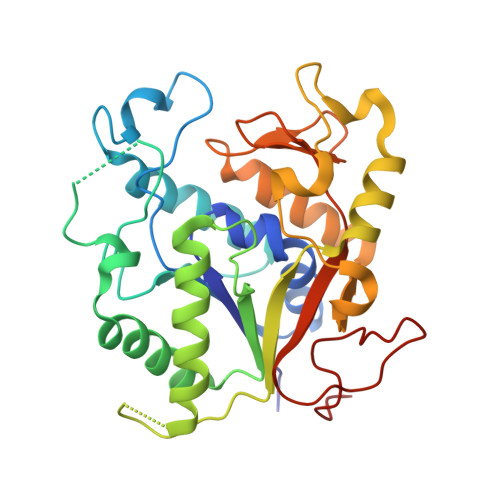
Mycothiol (1-D-myo-inosityl 2-(N-acetyl-L-cysteinyl)amido-2-deoxy-alpha-D-glucopyranoside, MSH or AcCys-GlcN-inositol (Ins)) is the major reducing agent in actinomycetes, including Mycobacterium tuberculosis. The biosynthesis of MSH involves a deacetylase that removes the acetyl group from the precursor GlcNAc-Ins to yield GlcN-Ins. The deacetylase (MshB) corresponds to Rv1170 of M. tuberculosis with a molecular mass of 33,400 Da. MshB is a Zn2+ metalloprotein, and the deacetylase activity is completely dependent on the presence of a divalent metal cation. We have determined the x-ray crystallographic structure of MshB, which reveals a protein that folds in a manner resembling lactate dehydrogenase in the N-terminal domain and a C-terminal domain consisting of two beta-sheets and two alpha-helices. The zinc binding site is in the N-terminal domain occupying a position equivalent to that of the NAD+ co-factor of lactate dehydrogenase. The Zn2+ is 5 coordinate with 3 residues from MshB (His-13, Asp-16, His-147) and two water molecules. One water would be displaced upon binding of substrate (GlcNAc-Ins); the other is proposed as the nucleophilic water assisted by the general base carboxylate of Asp-15. In addition to the Zn2+ providing electrophilic assistance in the hydrolysis, His-144 imidazole could form a hydrogen bond to the oxyanion of the tetrahedral intermediate. The extensive sequence identity of MshB, the deacetylase, with mycothiol S-conjugate amidase, an amide hydrolase that mediates detoxification of mycothiol S-conjugate xenobiotics, has allowed us to construct a faithful model of the catalytic domain of mycothiol S-conjugate amidase based on the structure of MshB.
Organizational Affiliation:
Canadian Institutes of Health Research, Group in Protein Structure and Function, Department of Biochemistry, Faculty of Medicine, University of Alberta, Edmonton, Alberta T6G 2H7, Canada.