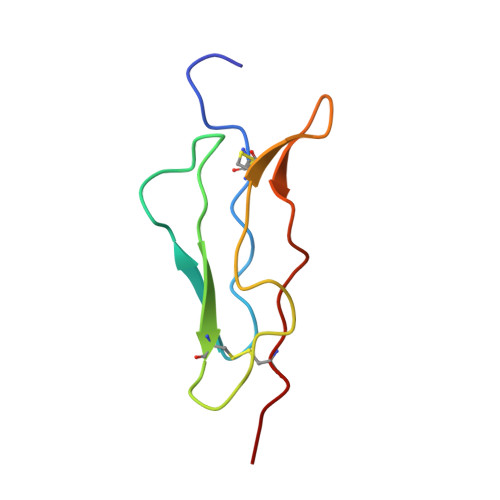
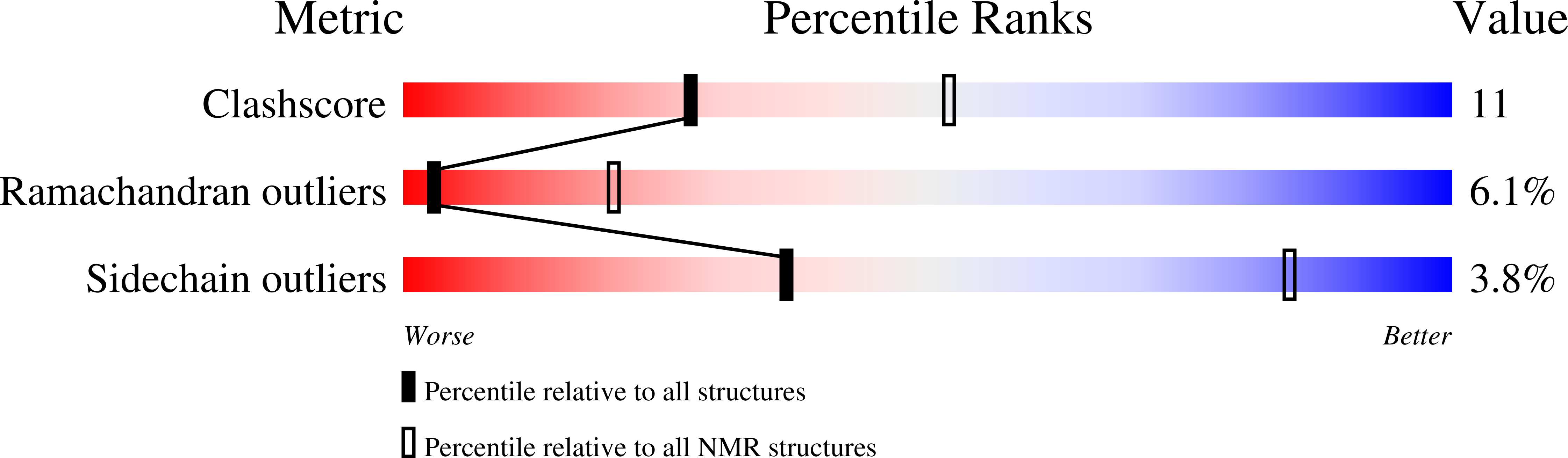Backbone dynamics of complement control protein (CCP) modules reveals mobility in binding surfaces.
O'Leary, J.M., Bromek, K., Black, G.M., Uhrinova, S., Schmitz, C., Wang, X., Krych, M., Atkinson, J.P., Uhrin, D., Barlow, P.N.(2004) Protein Sci 13: 1238-1250
- PubMed: 15096630
- DOI: https://doi.org/10.1110/ps.03582704
- Primary Citation of Related Structures:
1PPQ - PubMed Abstract:
The regulators of complement activation (RCA) are critical to health and disease because their role is to ensure that a complement-mediated immune response to infection is proportionate and targeted. Each protein contains an uninterrupted array of from four to 30 examples of the very widely occurring complement control protein (CCP, or sushi) module. The CCP modules mediate specific protein-protein and protein-carbohydrate interactions that are key to the biological function of the RCA and, paradoxically, provide binding sites for numerous pathogens. Although structural and mutagenesis studies of CCP modules have addressed some aspects of molecular recognition, there have been no studies of the role of molecular dynamics in the interaction of CCP modules with their binding partners. NMR has now been used in the first full characterization of the backbone dynamics of CCP modules. The dynamics of two individual modules-the 16th of the 30 modules of complement receptor type 1 (CD35), and the N-terminal module of membrane cofactor protein (CD46)-as well as their solution structures, are compared. Although both examples share broadly similar three-dimensional structures, many structurally equivalent residues exhibit different amplitudes and timescales of local backbone motion. In each case, however, regions of the module-surface implicated by mutagenesis as sites of interactions with other proteins include several mobile residues. This observation suggests further experiments to explore binding mechanisms and identify new binding sites.
Organizational Affiliation:
Schools of Chemistry and Biology, University of Edinburgh, Edinburgh EH9 3JJ, Scotland.