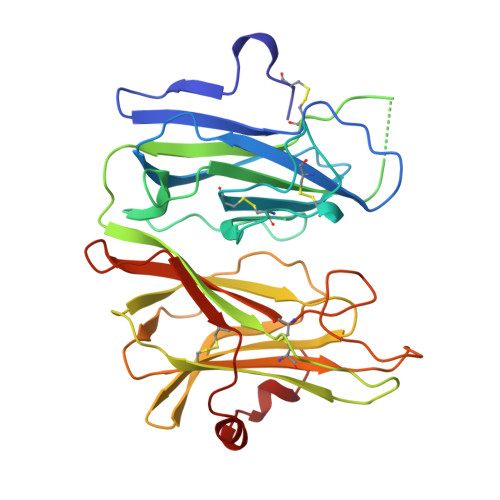
Amidation of bioactive peptides: the structure of peptidylglycine alpha-hydroxylating monooxygenase.
Prigge, S.T., Kolhekar, A.S., Eipper, B.A., Mains, R.E., Amzel, L.M.(1997) Science 278: 1300-1305
- PubMed: 9360928
- DOI: https://doi.org/10.1126/science.278.5341.1300
- Primary Citation of Related Structures:
1PHM - PubMed Abstract:
Many neuropeptides and peptide hormones require amidation at the carboxyl terminus for activity. Peptidylglycine alpha-amidating monooxygenase (PAM) catalyzes the amidation of these diverse physiological regulators. The amino-terminal domain of the bifunctional PAM protein is a peptidylglycine alpha-hydroxylating monooxygenase (PHM) with two coppers that cycle through cupric and cuprous oxidation states. The anomalous signal of the endogenous coppers was used to determine the structure of the catalytic core of oxidized rat PHM with and without bound peptide substrate. These structures strongly suggest that the PHM reaction proceeds via activation of substrate by a copper-bound oxygen species. The mechanistic and structural insight gained from the PHM structures can be directly extended to dopamine beta-monooxygenase.
Organizational Affiliation:
Department of Biophysics and Biophysical Chemistry, Johns Hopkins School of Medicine, Baltimore, MD 21205, USA.