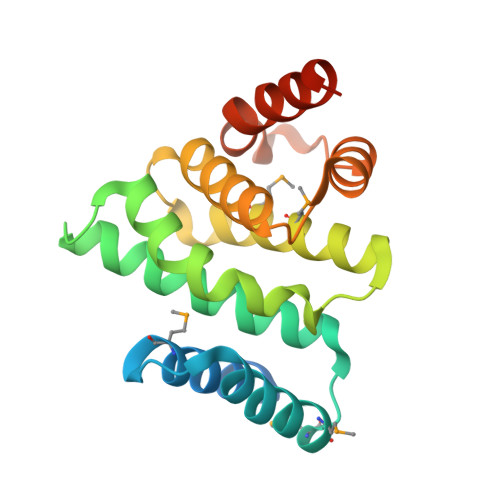
Structure of the catalytic fragment of translation initiation factor 2B and identification of a critically important catalytic residue.
Boesen, T., Mohammad, S.S., Pavitt, G.D., Andersen, G.R.(2004) J Biol Chem 279: 10584-10592
- PubMed: 14681227
- DOI: https://doi.org/10.1074/jbc.M311055200
- Primary Citation of Related Structures:
1PAQ - PubMed Abstract:
Eukaryotic initiation factor (eIF) 2B catalyzes the nucleotide activation of eIF2 to its active GTP-bound state. The exchange activity has been mapped to the C terminus of the eIF2Bepsilon subunit. We have determined the crystal structure of residues 544-704 from yeast eIF2Bepsilon at 2.3-A resolution, and this fragment is an all-helical protein built around the conserved aromatic acidic (AA) boxes also found in eIF4G and eIF5. The eight helices are organized in a manner similar to HEAT repeats. The molecule is highly asymmetric with respect to surface charge and conservation. One area in the N terminus is proposed to be directly involved in catalysis. In agreement with this hypothesis, mutation of glutamate 569 is shown to be lethal. An acidic belt and a second area in the C terminus containing residues from the AA boxes are important for binding to eIF2. Two mutations causing the fatal human genetic disease leukoencephalopathy with vanishing white matter are buried and appear to disrupt the structural integrity of the catalytic domain rather than interfering directly with catalysis or binding of eIF2.
Organizational Affiliation:
Department of Molecular Biology, Aarhus University, Denmark.