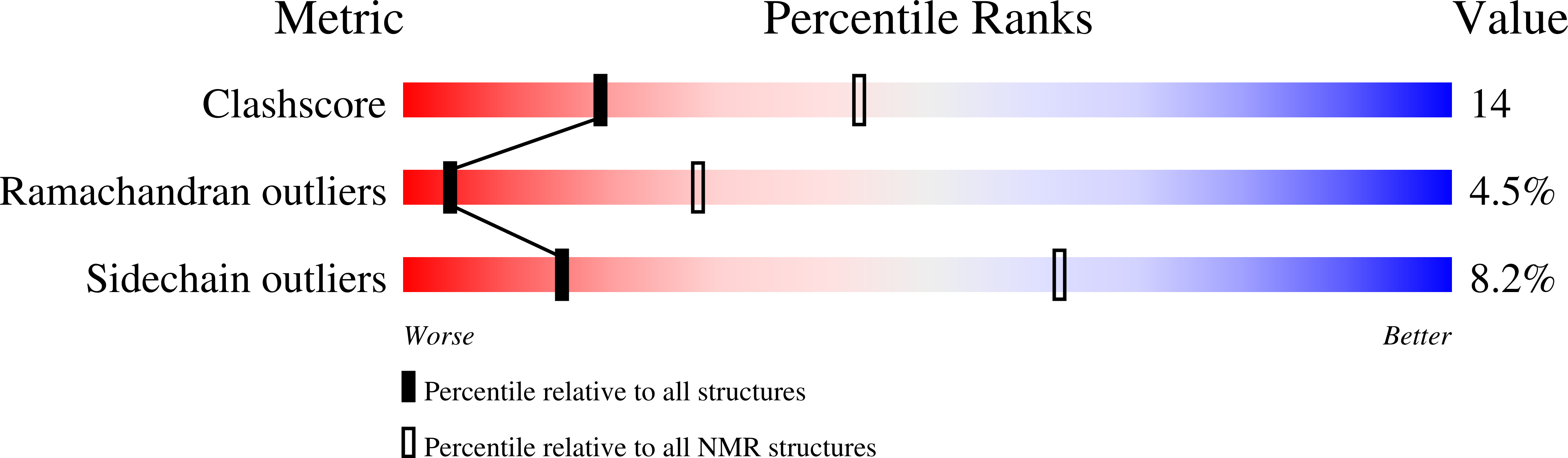Linearization of a naturally occurring circular protein maintains structure but eliminates hemolytic activity
Barry, D.G., Daly, N.L., Clark, R.J., Sando, L., Craik, D.J.(2003) Biochemistry 42: 6688-6695
- PubMed: 12779323
- DOI: https://doi.org/10.1021/bi027323n
- Primary Citation of Related Structures:
1ORX - PubMed Abstract:
Cyclotides are a recently discovered family of disulfide rich proteins from plants that contain a circular protein backbone. They are exceptionally stable, as exemplified by their use in native medicine of the prototypic cyclotide kalata B1. The peptide retains uterotonic activity after the plant from which it is derived is boiled to make a medicinal tea. The circular backbone is thought to be in part responsible for the stability of the cyclotides, and to investigate its role in determining structure and biological activity, an acyclic derivative, des-(24-28)-kalata B1, was chemically synthesized and purified. This derivative has five residues removed from the 29-amino acid circular backbone of kalata B1 in a loop region corresponding to a processing site in the biosynthetic precursor protein. Two-dimensional NMR spectra of the peptide were recorded, assigned, and used to identify a series of distance, angle, and hydrogen bonding restraints. These were in turn used to determine a representative family of solution structures. Of particular interest was a determination of the structural similarities and differences between des-(24-28)-kalata B1 and native kalata B1. Although the overall three-dimensional fold remains very similar to that of the native circular protein, removal of residues 24-28 of kalata B1 causes disruption of some structural features that are important to the overall stability. Furthermore, loss of hemolytic activity is associated with backbone truncation and linearization.
Organizational Affiliation:
Institute for Molecular Bioscience, Australian Research Council Special Research Centre for Functional and Applied Genomics, University of Queensland, Brisbane 4072, Australia.