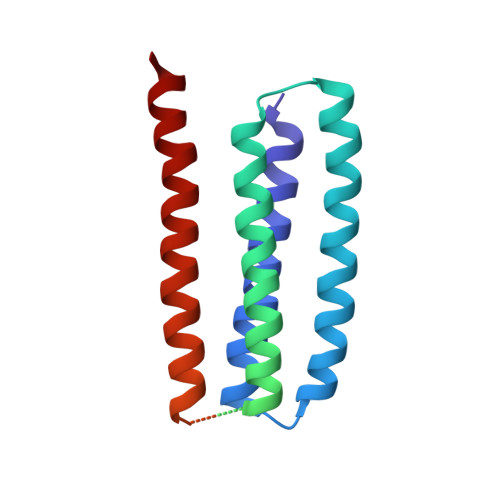
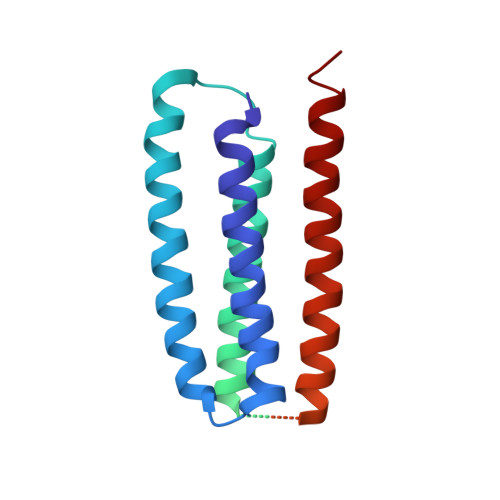
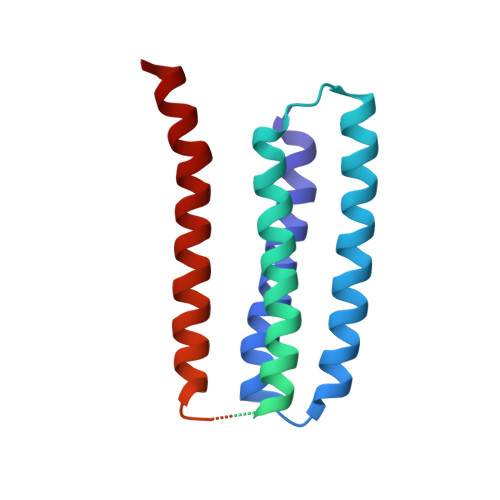
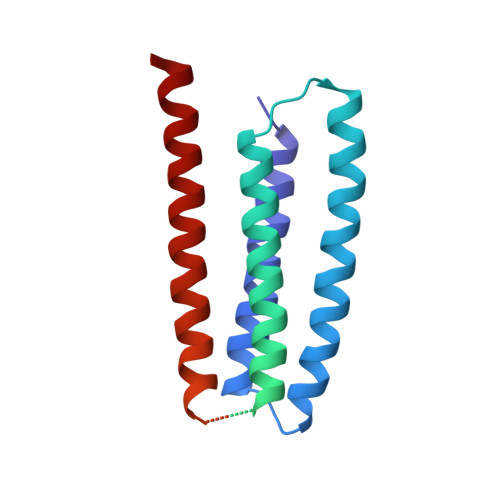
Structure and gating mechanism of the acetylcholine receptor pore.
Miyazawa, A., Fujiyoshi, Y., Unwin, N.(2003) Nature 423: 949-955
- PubMed: 12827192
- DOI: https://doi.org/10.1038/nature01748
- Primary Citation of Related Structures:
1OED - PubMed Abstract:
The nicotinic acetylcholine receptor controls electrical signalling between nerve and muscle cells by opening and closing a gated, membrane-spanning pore. Here we present an atomic model of the closed pore, obtained by electron microscopy of crystalline postsynaptic membranes. The pore is shaped by an inner ring of 5 alpha-helices, which curve radially to create a tapering path for the ions, and an outer ring of 15 alpha-helices, which coil around each other and shield the inner ring from the lipids. The gate is a constricting hydrophobic girdle at the middle of the lipid bilayer, formed by weak interactions between neighbouring inner helices. When acetylcholine enters the ligand-binding domain, it triggers rotations of the protein chains on opposite sides of the entrance to the pore. These rotations are communicated through the inner helices, and open the pore by breaking the girdle apart.
Organizational Affiliation:
RIKEN Harima Institute, 1-1-1 Kouto, Mikazuki-cho, Sayo, Hyogo 679-5148, Japan.