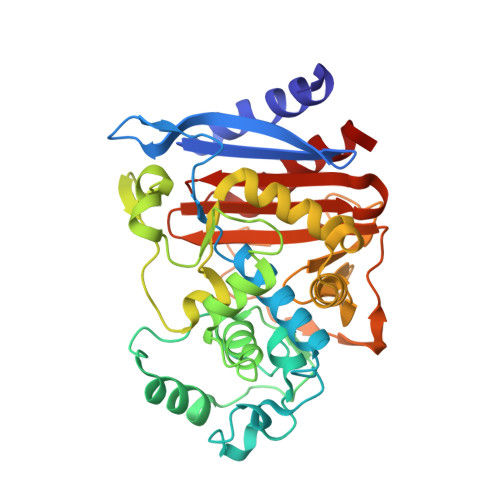
Structural Aspects for Evolution of beta-Lactamases from Penicillin-Binding Proteins
Meroueh, S.O., Minasov, G., Lee, W., Shoichet, B.K., Mobashery, S.(2003) J Am Chem Soc 125: 9612-9618
- PubMed: 12904027
- DOI: https://doi.org/10.1021/ja034861u
- Primary Citation of Related Structures:
1O07 - PubMed Abstract:
Penicillin-binding proteins (PBPs), biosynthetic enzymes of bacterial cell wall assembly, and beta-lactamases, resistance enzymes to beta-lactam antibiotics, are related to each other from an evolutionary point of view. Massova and Mobashery (Antimicrob. Agents Chemother. 1998, 42, 1-17) have proposed that for beta-lactamases to have become effective at their function as antibiotic resistance enzymes, they would have had to undergo structure alterations such that they would not interact with the peptidoglycan, which is the substrate for PBPs. A cephalosporin analogue, 7beta-[N-Acetyl-L-alanyl-gamma-D-glutamyl-L-lysine]-3-acetoxymethyl-3-cephem-carboxylic acid (compound 6), was conceived and synthesized to test this notion. The X-ray structure of the complex of this cephalosporin bound to the active site of the deacylation-deficient Q120L/Y150E variant of the class C AmpC beta-lactamase from Escherichia coli was solved at 1.71 A resolution. This complex revealed that the surface for interaction with the strand of peptidoglycan that acylates the active site, which is present in PBPs, is absent in the -lactamase active site. Furthermore, insertion of a peptide in the beta-lactamase active site at a location where the second strand of peptidoglycan in some PBPs binds has effectively abolished the possibility for such interaction with the beta-lactamase. A 2.6 ns dynamics simulation was carried out for the complex, which revealed that the peptidoglycan surrogate (i.e., the active-site-bound ligand) undergoes substantial motion and is not stabilized for binding within the active site. These factors taken together disclose the set of structure modifications in the antibiotic resistance enzyme that prevent it from interacting with the peptidoglycan, en route to achieving catalytic proficiency for their intended function.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of Notre Dame, Notre Dame, Indiana 46556, USA.