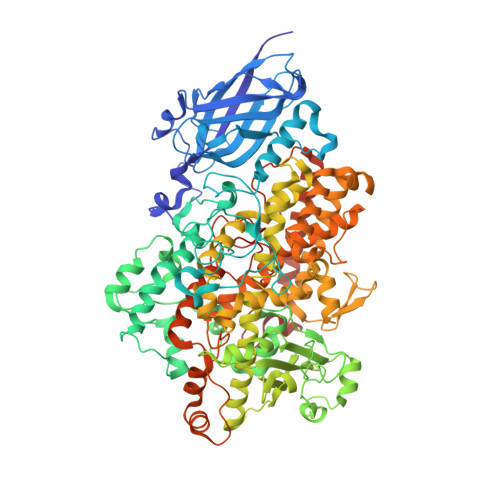
Soybean lipoxygenase-3 in complex with 4-nitrocatechol.
Skrzypczak-Jankun, E., Borbulevych, O.Y., Jankun, J.(2004) Acta Crystallogr D Biol Crystallogr 60: 613-615
- PubMed: 14993710
- DOI: https://doi.org/10.1107/S0907444904000861
- Primary Citation of Related Structures:
1NO3 - PubMed Abstract:
4-Nitrocatechol (4NC) is a known inhibitor of lipoxygenase. This work presents the X-ray structure of soybean lipoxygenase-3 in complex with 4NC refined at 2.15 A resolution. The X-ray analysis shows 4NC near iron with partial occupancy, blocking access to Fe but not covalently bound to it. The two hydroxyl groups interact with the C-terminus (4-OH) and His523 ND1 (3-OH). The residues surrounding the iron cofactor, His518*, His523, His709, Ile857* COO(-) and water, form a trigonal bipyramid with the residues marked with asterisks in the axial positions. The water bound to iron and the presence of the inhibitor seem to be responsible for the rearrangements and changes in the geometry of the ligand distribution and confirm the displacement of His518 from iron coordination. A description of the catechol binding contributes to the understanding of lipoxygenase inhibition and the participation of its co-oxidative activity in the utilization of natural flavonoids.
Organizational Affiliation:
Medical College of Ohio, Urology Research Center, Department of Urology, 3065 Arlington Avenue, Toledo OH 43614, USA. eskrzypczak@mco.edu