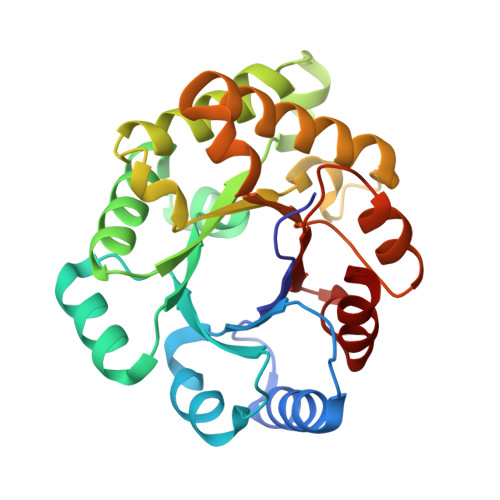
Three new crystal structures of point mutation variants of monoTIM: conformational flexibility of loop-1, loop-4 and loop-8.
Borchert, T.V., Kishan, K.V., Zeelen, J.P., Schliebs, W., Thanki, N., Abagyan, R., Jaenicke, R., Wierenga, R.K.(1995) Structure 3: 669-679
- PubMed: 8591044
- DOI: https://doi.org/10.1016/s0969-2126(01)00202-7
- Primary Citation of Related Structures:
1MSS, 1TTI, 1TTJ - PubMed Abstract:
Wild-type triosephosphate isomerase (TIM) is a very stable dimeric enzyme. This dimer can be converted into a stable monomeric protein (monoTIM) by replacing the 15-residue interface loop (loop-3) by a shorter, 8-residue, loop. The crystal structure of monoTIM shows that two active-site loops (loop-1 and loop-4), which are at the dimer interface in wild-type TIM, have acquired rather different structural properties. Nevertheless, monoTIM has residual catalytic activity.
Organizational Affiliation:
EMBL, Heidelberg, Germany.