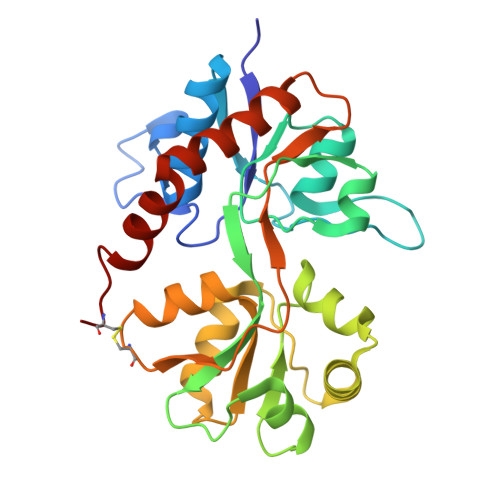
Structural basis for partial agonist action at ionotropic glutamate receptors
Jin, R., Banke, T.G., Mayer, M.L., Traynelis, S.F., Gouaux, E.(2003) Nat Neurosci 6: 803-810
- PubMed: 12872125
- DOI: https://doi.org/10.1038/nn1091
- Primary Citation of Related Structures:
1MQG, 1MQH, 1MQI, 1MQJ - PubMed Abstract:
An unresolved problem in understanding neurotransmitter receptor function concerns the mechanism(s) by which full and partial agonists elicit different amplitude responses at equal receptor occupancy. The widely held view of 'partial agonism' posits that resting and active states of the receptor are in equilibrium, and partial agonists simply do not shift the equilibrium toward the active state as efficaciously as full agonists. Here we report findings from crystallographic and electrophysiological studies of the mechanism of activation of an AMPA-subtype glutamate receptor ion channel. In these experiments, we used 5-substituted willardiines, a series of partial agonists that differ by only a single atom. Our results show that the GluR2 ligand-binding core can adopt a range of ligand-dependent conformational states, which in turn control the open probability of discrete subconductance states of the intact ion channel. Our findings thus provide a structure-based model of partial agonism.
Organizational Affiliation:
Department of Biochemistry and Molecular Biophysics, Columbia University, 650 West 168 Street, New York, New York 10032, USA.