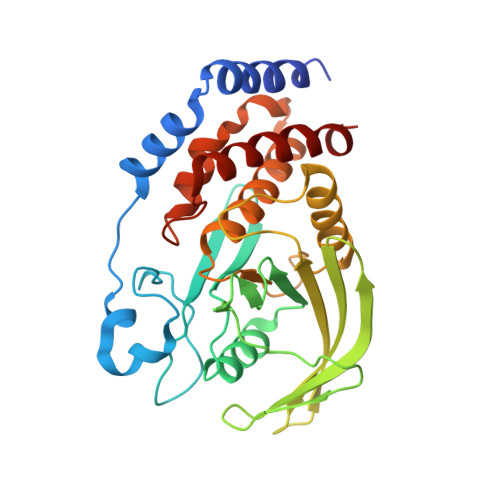
The structure of PTP-1B in complex with a peptide inhibitor reveals an alternative binding mode for bisphosphonates.
Asante-Appiah, E., Patel, S., Dufresne, C., Roy, P., Wang, Q., Patel, V., Friesen, R.W., Ramachandran, C., Becker, J.W., Leblanc, Y., Kennedy, B.P., Scapin, G.(2002) Biochemistry 41: 9043-9051
- PubMed: 12119018
- DOI: https://doi.org/10.1021/bi0259554
- Primary Citation of Related Structures:
1LQF - PubMed Abstract:
Inhibitors of PTP-1B could be therapeutically beneficial in the treatment of type 2 diabetes. Owing to the large number of phosphatases in the cell, inhibitors against PTP-1B must not only be potent but selective as well. N-Benzoyl-L-glutamyl-[4-phosphono(difluoromethyl)]-L-phenylalanine-[4-phosphono(difluoro-methyl)]-L-phenylalanineamide (BzN-EJJ-amide) is a low nanomolar inhibitor of PTP-1B that shows selectivity over several protein tyrosine phosphatases. To gain an insight into the basis of its potency and selectivity, we evaluated several analogues of the inhibitor and introduced amino acid substitutions into PTP-1B by site-directed mutagenesis. We also determined the crystal structure of PTP-1B in complex with BzN-EJJ-amide at 2.5 A resolution. Our results indicate that the high inhibitory potency is due to interactions of several of its chemical groups with specific protein residues. An interaction between BzN-EJJ-amide and Asp48 is of particular significance, as substitution of Asp48 to alanine resulted in a 100-fold loss in potency. The crystal structure also revealed an unexpected binding orientation for a bisphosphonate inhibitor on PTP-1B, where the second difluorophosphonomethyl phenylalanine (F(2)PMP) moiety is bound close to Arg47 rather than in the previously identified second aryl phosphate site demarked by Arg24 and Arg254. Our results suggest that potent and selective PTP-1B inhibitors may be designed by targeting the region containing Arg47 and Asp48.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Merck Frosst Center for Therapeutic Research, Pointe-Claire - Dorval H9R 4P8, Canada. Ernest_Asanteappiah@Merck.com