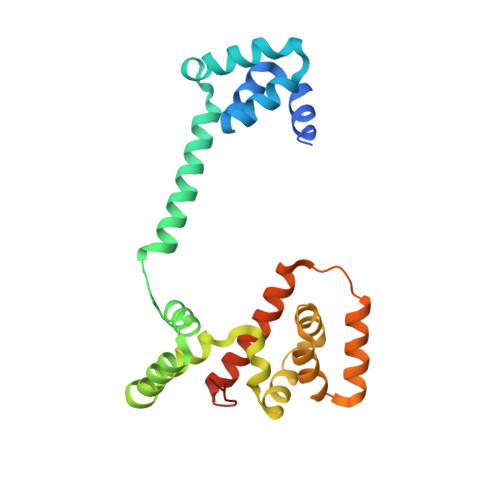
Crystal structure of the middle and C-terminal domains of the flagellar rotor protein FliG.
Brown, P.N., Hill, C.P., Blair, D.F.(2002) EMBO J 21: 3225-3234
- PubMed: 12093724
- DOI: https://doi.org/10.1093/emboj/cdf332
- Primary Citation of Related Structures:
1LKV - PubMed Abstract:
The FliG protein is essential for assembly, rotation and clockwise/counter-clockwise (CW/CCW) switching of the bacterial flagellum. About 25 copies of FliG are present in a large rotor-mounted assembly termed the 'switch complex', which also contains the proteins FliM and FliN. Mutational studies have identified the segments of FliG most crucial for flagellar assembly, rotation and switching. The structure of the C-terminal domain, which functions specifically in rotation, was reported previously. Here, we describe the crystal structure of a larger fragment of the FliG protein from Thermotoga maritima, which encompasses the middle and C-terminal parts of the protein (termed FliG-MC). The FliG-MC molecule consists of two compact globular domains, linked by an alpha-helix and an extended segment that contains a well-conserved Gly-Gly motif. Mutational studies indicate that FliM binds to both of the globular domains, and given the flexibility of the linking segment, FliM is likely to determine the relative orientation of the domains in the flagellum. We propose a model for the organization of FliG-MC molecules in the flagellum, and suggest that CW/CCW switching might occur by movement of the C-terminal domain relative to other parts of FliG, under the control of FliM.
Organizational Affiliation:
Department of Biology, University of Utah, Salt Lake City, UT, USA.