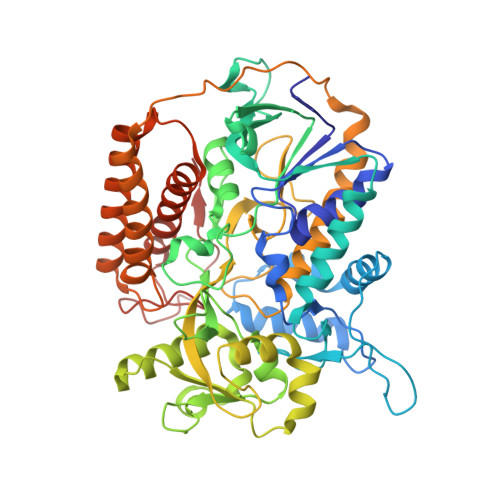
Structure of FAD-bound L-aspartate oxidase: insight into substrate specificity and catalysis.
Bossi, R.T., Negri, A., Tedeschi, G., Mattevi, A.(2002) Biochemistry 41: 3018-3024
- PubMed: 11863440
- DOI: https://doi.org/10.1021/bi015939r
- Primary Citation of Related Structures:
1KNP, 1KNR - PubMed Abstract:
L-Aspartate oxidase (Laspo) catalyzes the conversion of L-Asp to iminoaspartate, the first step in the de novo biosynthesis of NAD(+). This bacterial pathway represents a potential drug target since it is absent in mammals. The Laspo R386L mutant was crystallized in the FAD-bound catalytically competent form and its three-dimensional structure determined at 2.5 A resolution in both the native state and in complex with succinate. Comparison of the R386L holoprotein with the wild-type apoenzyme [Mattevi, A., Tedeschi, G., Bacchella, L., Coda, A., Negri, A., and Ronchi, S. (1999) Structure 7, 745-756] reveals that cofactor incorporation leads to the ordering of two polypeptide segments (residues 44-53 and 104-141) and to a 27 degree rotation of the capping domain. This motion results in the formation of the active site cavity, located at the interface between the capping domain and the FAD-binding domain. The structure of the succinate complex indicates that the cavity surface is decorated by two clusters of H-bond donors that anchor the ligand carboxylates. Moreover, Glu121, which is strictly conserved among Laspo sequences, is positioned to interact with the L-Asp alpha-amino group. The architecture of the active site of the Laspo holoenzyme is remarkably similar to that of respiratory fumarate reductases, providing strong evidence for a common mechanism of catalysis in Laspo and flavoproteins of the succinate dehydrogenase/fumarate reductase family. This implies that Laspo is mechanistically distinct from other flavin-dependent amino acid oxidases, such as the prototypical D-amino acid oxidase.
Organizational Affiliation:
Dipartimento di Genetica e Microbiologia, Università di Pavia, via Abbiategrasso 207, 27100 Pavia, Italy.