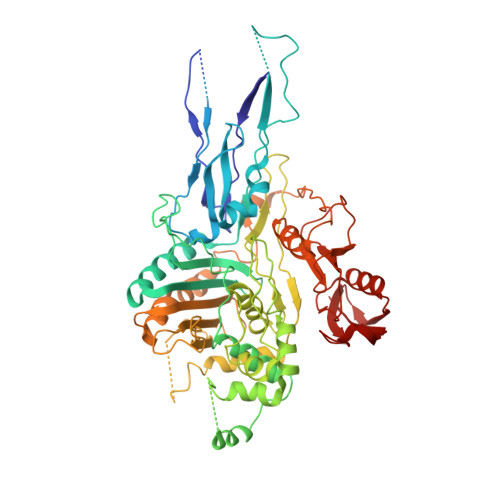
Crystal structure of PBP2x from a highly penicillin-resistant Streptococcus pneumoniae clinical isolate: a mosaic framework containing 83 mutations.
Dessen, A., Mouz, N., Gordon, E., Hopkins, J., Dideberg, O.(2001) J Biol Chem 276: 45106-45112
- PubMed: 11553637
- DOI: https://doi.org/10.1074/jbc.M107608200
- Primary Citation of Related Structures:
1K25 - PubMed Abstract:
Penicillin-binding proteins (PBPs) are the main targets for beta-lactam antibiotics, such as penicillins and cephalosporins, in a wide range of bacterial species. In some Gram-positive strains, the surge of resistance to treatment with beta-lactams is primarily the result of the proliferation of mosaic PBP-encoding genes, which encode novel proteins by recombination. PBP2x is a primary resistance determinant in Streptococcus pneumoniae, and its modification is an essential step in the development of high level beta-lactam resistance. To understand such a resistance mechanism at an atomic level, we have solved the x-ray crystal structure of PBP2x from a highly penicillin-resistant clinical isolate of S. pneumoniae, Sp328, which harbors 83 mutations in the soluble region. In the proximity of the Sp328 PBP2x* active site, the Thr(338) --> Ala mutation weakens the local hydrogen bonding network, thus abrogating the stabilization of a crucial buried water molecule. In addition, the Ser(389) --> Leu and Asn(514) --> His mutations produce a destabilizing effect that generates an "open" active site. It has been suggested that peptidoglycan substrates for beta-lactam-resistant PBPs contain a large amount of abnormal, branched peptides, whereas sensitive strains tend to catalyze cross-linking of linear forms. Thus, in vivo, an "open" active site could facilitate the recognition of distinct, branched physiological substrates.
Organizational Affiliation:
Laboratoire de Cristallographie Macromoléculaire, Institut de Biologie Structurale Jean-Pierre Ebel (CNRS/Commissariat à l'Energie Atomique), 41, rue Jules Horowitz, 38027 Grenoble, France. dessen@ibs.fr