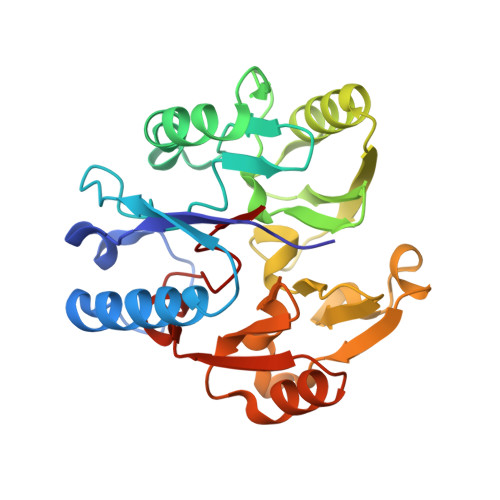
Structural insights into the hydrolysis of cellular nitric oxide synthase inhibitors by dimethylarginine dimethylaminohydrolase.
Murray-Rust, J., Leiper, J., McAlister, M., Phelan, J., Tilley, S., Santa Maria, J., Vallance, P., McDonald, N.(2001) Nat Struct Biol 8: 679-683
- PubMed: 11473257
- DOI: https://doi.org/10.1038/90387
- Primary Citation of Related Structures:
1H70 - PubMed Abstract:
Nitric oxide synthase is inhibited by asymmetric NG-methylated derivatives of arginine whose cellular levels are controlled in part by dimethylarginine dimethylaminohydrolase (DDAH, EC 3.5.3.18). Levels of asymmetric NG,NG-dimethylarginine (ADMA) are known to correlate with certain disease states. Here, the first structure of a DDAH shows an unexpected similarity to arginine:glycine amidinotransferase (EC 2.1.4.1) and arginine deiminase (EC 3.5.3.6), thus defining a superfamily of arginine-modifying enzymes. The identification of a Cys-His-Glu catalytic triad and the structures of a Cys to Ser point mutant bound to both substrate and product suggest a reaction mechanism. Comparison of the ADMA-DDAH and arginine-amidinotransferase complexes reveals a dramatic rotation of the substrate that effectively maintains the orientation of the scissile bond of the substrate with respect to the catalytic residues. The DDAH structure will form a basis for the rational design of selective inhibitors, which are of potential use in modulating NO synthase activity in pathological settings.
Organizational Affiliation:
School of Crystallography, Birkbeck, Malet Street, London WC1E 7HX, UK.