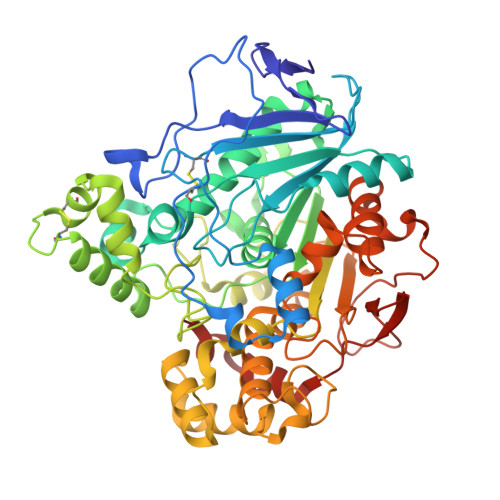
Structural Insights Into the Lipase/Esterase Behavior in the Candida Rugosa Lipases Family: Crystal Structure of the Lipase 2 Isoenzyme at 1.97A Resolution
Mancheno, J.M., Pernas, M.A., Martinez, M.J., Ochoa, B., Rua, M.L., Hermoso, J.A.(2003) J Mol Biol 332: 1059
- PubMed: 14499609
- DOI: https://doi.org/10.1016/j.jmb.2003.08.005
- Primary Citation of Related Structures:
1GZ7 - PubMed Abstract:
The yeast Candida rugosa produces several closely related extracellular lipases that differ in their substrate specificity. Here, we report the crystal structure of the isoenzyme lipase 2 at 1.97A resolution in its closed conformation. Lipase 2 shows a 79.4% amino acid sequence identity with lipase 1 and 82.2% with lipase 3, which makes it relevant to compare these three isoenzymes. Despite this high level of sequence identity, structural comparisons reveal several amino acid changes affecting the flap (residue 69), the substrate-binding pocket (residues 127, 132 and 450) and the mouth of the hydrophobic tunnel (residues 296 and 344), which may be responsible for the different substrate specificity and catalytic properties of this group of enzymes. Also, these comparisons reveal two distinct regions in the hydrophobic tunnel: a phenylalanyl-rich region and an aliphatic-rich region. Whereas this last region is essentially identical in the three isoenzymes, the phenylalanyl content in the first one is specific for each lipase, resulting in a different environment of the catalytic triad residues, which probably tunes finely their lipase/esterase character. The greater structural similarity observed between the monomeric form of lipase 3 and lipase 2 concerning the above-mentioned key residues led us to propose a significant esterase activity for this last protein. This enzymatic activity has been confirmed with biochemical experiments using cholesteryl [1-14C]oleate as substrate. Surprisingly, lipase 2 is a more efficient esterase than lipase 3, showing a twofold specific activity against cholesteryl [1-14C]oleate in our experimental conditions. These results show that subtle amino acid changes within a highly conserved protein fold may produce protein variants endowed with new enzymatic properties.
Organizational Affiliation:
Grupo de Cristalografía Macromolecular y Biología Estructural, Instituto Química-Física Rocasolano C.S.I.C., Serrano 119, 28006, Madrid, Spain.