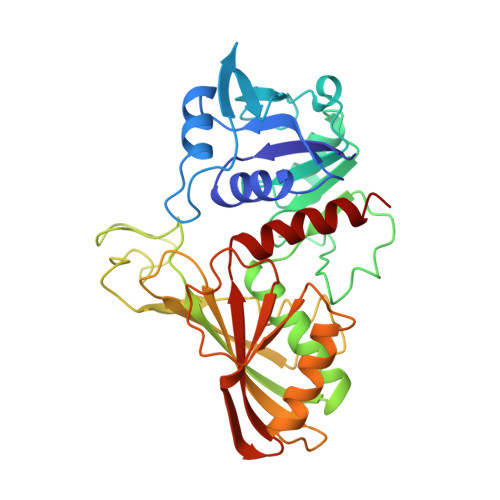
Comparison of the structures of wild-type and a N313T mutant of Escherichia coli glyceraldehyde 3-phosphate dehydrogenases: implication for NAD binding and cooperativity.
Duee, E., Olivier-Deyris, L., Fanchon, E., Corbier, C., Branlant, G., Dideberg, O.(1996) J Mol Biol 257: 814-838
- PubMed: 8636984
- DOI: https://doi.org/10.1006/jmbi.1996.0204
- Primary Citation of Related Structures:
1GAD, 1GAE - PubMed Abstract:
The crystal structure of wild-type and N313T mutant glyceraldehyde 3-phosphate dehydrogenases from Escherichia coli was determined in the presence of NAD at 1.8 angstrom and 2.17 angstrom, respectively. The structure of the monomer and of the tetramer are similar to those observed for other GAPDHs. An exhaustive analysis of the hydrophobic clusters and the hydrogen bond networks explain the high degree of sequence conservation in GAPDHs. The structural effect of the N313T mutation is a change in the (phi,psi) angles of nearby residues Asn236 and Val237, while the structure around the mutated residue remains unchanged. A detailed comparison of the wild-type and N313T mutant E. coli GAPDH with the apo and holo forms of Bacillus stearothermophilus GAPDH is carried out in relation to the apo --> holo transition. An unbiased set of about 60 residues, whose C(alpha) atoms remain in the same relative position in the different forms of the tetramer, is defined as the tetramer "core" which acts as a fixed scaffold around which structural rearrangements occur during the apo --> holo transition. This core essentially includes beta-strands from the beta-sheets forming the O-P and Q-R interfaces, in particular strand beta1 which bears catalytic residue His176. During the apo --> holo transition, dimer O-P rotates around the molecular P-axis by about +1 degrees, and dimer O-R by about -1 degrees. Further rotations of the NAD binding domain relative to the catalytic domain are discussed in relation to the molecular symmetry. The possible effect on NAD binding cooperativity of mutations around the tetramer core is exemplified by residue 252. The presence of a conserved hydrophilic patch embedded in the hydrophobic O-P interface is highlighted. A mechanism for substrate binding, different from those currently proposed, is described where the hydroxyl group of the substrate C(2) atom is hydrogen bonded to Cys149N.
Organizational Affiliation:
Laboratoire de Cristallographie Macromoléculaire, Institut de Biologie Structurale Jean-Pierre EBEL, Grenoble, France.