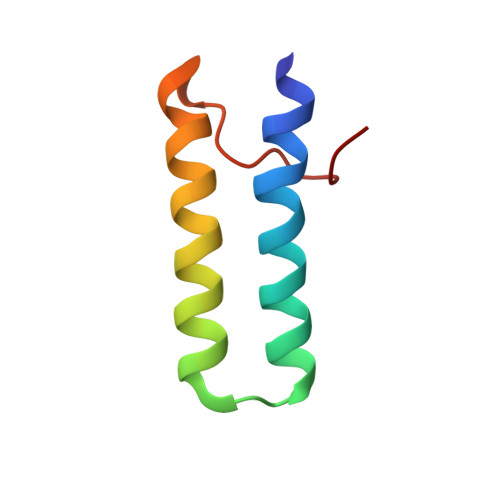
Structural analysis of multifunctional peptide motifs in human bifunctional tRNA synthetase: identification of RNA-binding residues and functional implications for tandem repeats.
Jeong, E.J., Hwang, G.S., Kim, K.H., Kim, M.J., Kim, S., Kim, K.S.(2000) Biochemistry 39: 15775-15782
- PubMed: 11123902
- DOI: https://doi.org/10.1021/bi001393h
- Primary Citation of Related Structures:
1FYJ - PubMed Abstract:
Human bifunctional glutamyl-prolyl-tRNA synthetase (EPRS) contains three tandem repeats linking the two catalytic domains. These repeated motifs have been shown to be involved in protein-protein and protein-nucleic acid interactions. The single copy of the homologous motifs has also been found in several different aminoacyl-tRNA synthetases. The solution structure of repeat 1 (EPRS-R1) and the secondary structure of the whole appended domain containing three repeated motifs in EPRS (EPRS-R123) was determined by nuclear magnetic resonance (NMR) spectroscopy. EPRS-R1 consists of two helices (residues 679-699 and 702-721) arranged in a helix-turn-helix, which is similar to other RNA binding proteins and the j-domain of DnaJ, and EPRS-R123 is composed of three helix-turn-helix motifs linked by an unstructured loop. When tRNA is bound to the appended domain, chemical shifts of several residues in each repeat are perturbed. However, the perturbed residues in each repeat are not the same although they are in the same binding surface, suggesting that each repeat in the appended domain is dynamically arranged to maximize contacts with tRNA. The affinity of tRNA to the three-repeated motif was much higher than to the single motif. These results indicate that each of the repeated motifs has a weak intrinsic affinity for tRNA, but the repetition of the motifs may be required to enhance binding affinity. Thus, the results of this work gave information on the RNA-binding mode of the multifunctional peptide motif attached to different ARSs and the functional reason for the repetition of this motif.
Organizational Affiliation:
Structural Biology Center, Korea Institute of Science and Technology (KIST), Cheongryang Box 131, Seoul, 130-650, Korea.