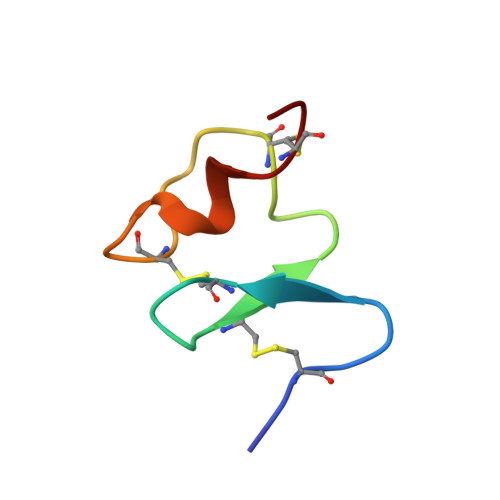
Three-dimensional NMR structure of the sixth ligand-binding module of the human LDL receptor: comparison of two adjacent modules with different ligand binding specificities.
Clayton, D., Brereton, I.M., Kroon, P.A., Smith, R.(2000) FEBS Lett 479: 118-122
- PubMed: 10981718
- DOI: https://doi.org/10.1016/s0014-5793(00)01842-1
- Primary Citation of Related Structures:
1F8Z - PubMed Abstract:
The sixth ligand-binding module of the low-density lipoprotein receptor contributes to the binding of apolipoprotein B100-containing lipoproteins. 1H NMR spectroscopy, DYANA and X-PLOR structure calculations were used to determine that this module has a well defined structure with a backbone conformation similar to other modules. Structures from calculations that simulated the presence of a calcium ion showed increased resolution without large increases in energy, increased deviations from idealised geometry or violations of experimental constraints. Investigation of the surface properties of this module indicates there are significant differences from the fifth module, which binds apolipoprotein E-containing lipoproteins in addition to apolipoprotein B100-containing lipoproteins.
Organizational Affiliation:
Department of Biochemistry, University of Queensland, Brisbane, Australia.