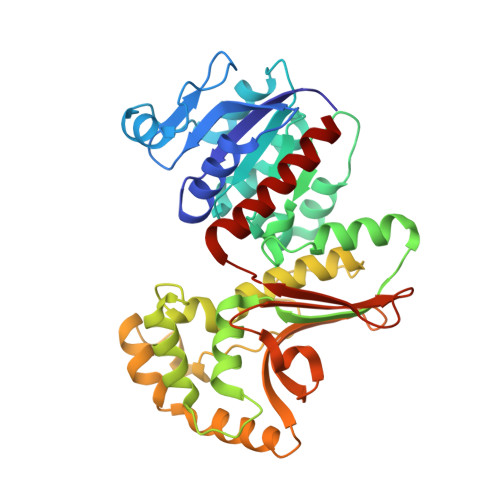
Crystal structures of homoserine dehydrogenase suggest a novel catalytic mechanism for oxidoreductases.
DeLaBarre, B., Thompson, P.R., Wright, G.D., Berghuis, A.M.(2000) Nat Struct Biol 7: 238-244
- PubMed: 10700284
- DOI: https://doi.org/10.1038/73359
- Primary Citation of Related Structures:
1EBF, 1EBU - PubMed Abstract:
The structure of the antifungal drug target homoserine dehydrogenase (HSD) was determined from Saccharomyces cerevisiae in apo and holo forms, and as a ternary complex with bound products, by X-ray diffraction. The three forms show that the enzyme is a dimer, with each monomer composed of three regions, the nucleotide-binding region, the dimerization region and the catalytic region. The dimerization and catalytic regions have novel folds, whereas the fold of the nucleotide-binding region is a variation on the Rossmann fold. The novel folds impose a novel composition and arrangement of active site residues when compared to all other currently known oxidoreductases. This observation, in conjunction with site-directed mutagenesis of active site residues and steady-state kinetic measurements, suggest that HSD exhibits a new variation on dehydrogenase chemistry.
Organizational Affiliation:
Antimicrobial Research Centre and Department of Biochemistry, McMaster University, 1200 Main Street West, Hamilton, Ontario, Canada L8N 3Z5.