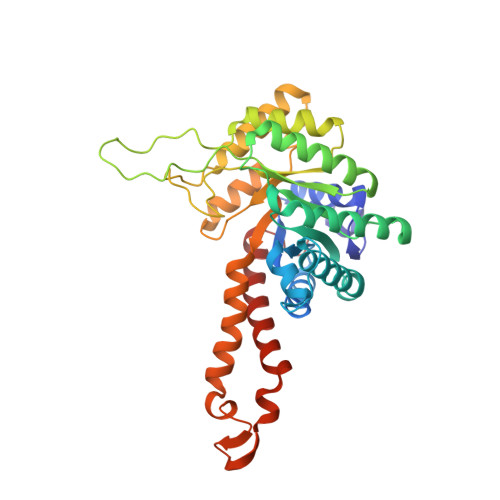
Novel active site in Escherichia coli fructose 1,6-bisphosphate aldolase.
Blom, N.S., Tetreault, S., Coulombe, R., Sygusch, J.(1996) Nat Struct Biol 3: 856-862
- PubMed: 8836102
- DOI: https://doi.org/10.1038/nsb1096-856
- Primary Citation of Related Structures:
1DOS - PubMed Abstract:
The molecular architecture of the Class II E. coli fructose 1,6-bisphosphate aldolase dimer was determined to 1.6 A resolution. The subunit fold corresponds to a singly wound alpha/beta-barrel with an active site located on the beta-barrel carboxyl side of each subunit. In each subunit there are two mutually exclusive zinc metal ion binding sites, 3.2 A apart; the exclusivity is mediated by a conformational transition involving side-chain rotations by chelating histidine residues. A binding site for K+ and NH4+ activators was found near the beta-barrel centre. Although Class I and Class II aldolases catalyse identical reactions, their active sites do not share common amino acid residues, are structurally dissimilar, and from sequence comparisons appear to be evolutionary distinct.
Organizational Affiliation:
Départment de biochimie, Université de Montréal, Canada.