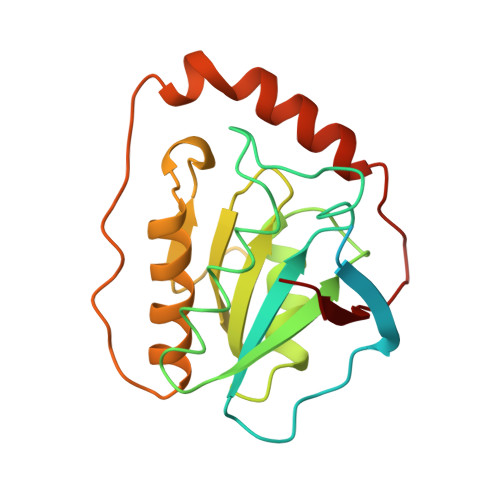
A new mode of B12 binding and the direct participation of a potassium ion in enzyme catalysis: X-ray structure of diol dehydratase.
Shibata, N., Masuda, J., Tobimatsu, T., Toraya, T., Suto, K., Morimoto, Y., Yasuoka, N.(1999) Structure 7: 997-1008
- PubMed: 10467140
- DOI: https://doi.org/10.1016/s0969-2126(99)80126-9
- Primary Citation of Related Structures:
1DIO - PubMed Abstract:
Diol dehydratase is an enzyme that catalyzes the adenosylcobalamin (coenzyme B12) dependent conversion of 1,2-diols to the corresponding aldehydes. The reaction initiated by homolytic cleavage of the cobalt-carbon bond of the coenzyme proceeds by a radical mechanism. The enzyme is an alpha2beta2gamma2 heterooligomer and has an absolute requirement for a potassium ion for catalytic activity. The crystal structure analysis of a diol dehydratase-cyanocobalamin complex was carried out in order to help understand the mechanism of action of this enzyme.
Organizational Affiliation:
Department of Life Science, Himeji Institute of Technology, Hyogo, Japan.