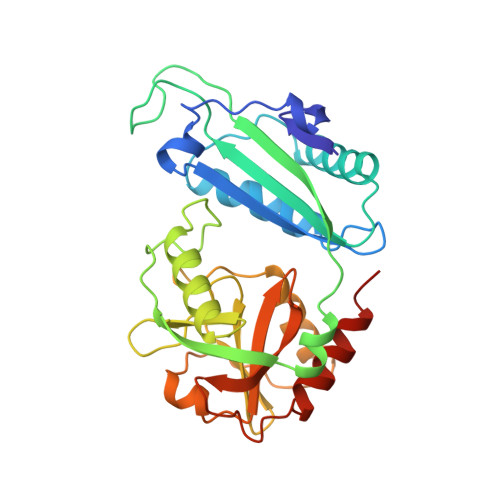
Crystal structure of a D-amino acid aminotransferase: how the protein controls stereoselectivity.
Sugio, S., Petsko, G.A., Manning, J.M., Soda, K., Ringe, D.(1995) Biochemistry 34: 9661-9669
- PubMed: 7626635
- DOI: https://doi.org/10.1021/bi00030a002
- Primary Citation of Related Structures:
1DAA - PubMed Abstract:
The three-dimensional structure of D-amino acid aminotransferase (D-AAT) in the pyridoxamine phosphate form has been determined crystallographically. The fold of this pyridoxal phosphate (PLP)-containing enzyme is completely different from those of any of the other enzymes that utilize PLP as part of their mechanism and whose structures are known. However, there are some striking similarities between the active sites of D-AAT and the corresponding enzyme that transaminates L-amino acids, L-aspartate aminotransferase. These similarities represent convergent evolution to a common solution of the problem of enforcing transamination chemistry on the PLP cofactor. Implications of these similarities are discussed in terms of their possible roles in the stabilization of intermediates of a transamination reaction. In addition, sequence similarity between D-AAT and branched chain L-amino acid aminotransferase suggests that this latter enzyme will also have a fold similar to that of D-AAT.
Organizational Affiliation:
Department of Biochemistry, Brandeis University, Waltham, Massachusetts 02254, USA.