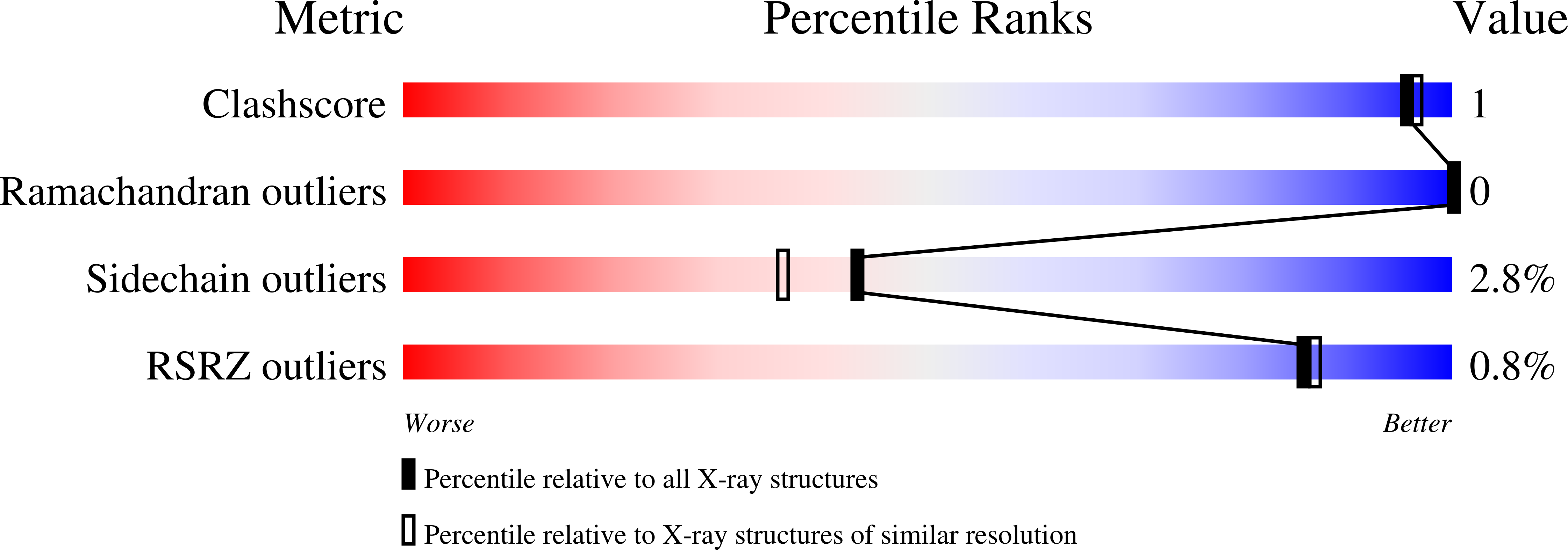
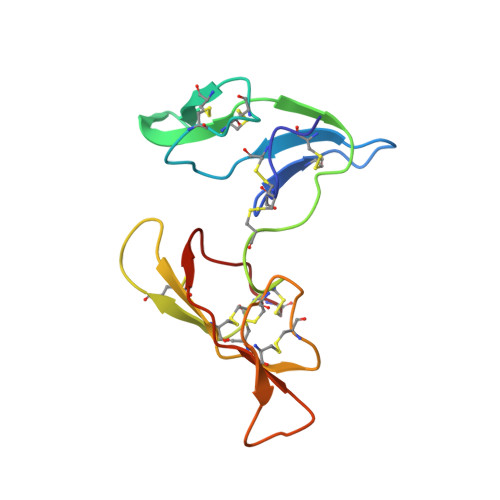
Crystal structure of a 16 kDa double-headed Bowman-Birk trypsin inhibitor from barley seeds at 1.9 A resolution.
Song, H.K., Kim, Y.S., Yang, J.K., Moon, J., Lee, J.Y., Suh, S.W.(1999) J Mol Biol 293: 1133-1144
- PubMed: 10547291
- DOI: https://doi.org/10.1006/jmbi.1999.3239
- Primary Citation of Related Structures:
1C2A - PubMed Abstract:
The Bowman-Birk trypsin inhibitor from barley seeds (BBBI) consists of 125 amino acid residues with two inhibitory loops. Its crystal structure in the free state has been determined by the multiwavelength anomalous diffraction (MAD) method and has been refined to a crystallographic R-value of 19.1 % for 8.0-1.9 A data. This is the first report on the structure of a 16 kDa double-headed Bowman-Birk inhibitor (BBI) from monocotyledonous plants and provides the highest resolution picture of a BBI to date. The BBBI structure consists of 11 beta-strands and the loops connecting these beta-strands but it lacks alpha-helices. BBBI folds into two compact domains of similar tertiary structure. Each domain shares the same overall fold with 8 kDa dicotyledonous BBIs. The five disulfide bridges in each domain are a subset of the seven disulfide bridges in 8 kDa dicotyledonous BBIs. Two buried water molecules form hydrogen bonds to backbone atoms in the core of each domain. One interesting feature of this two-domain inhibitor structure is that the two P1 residues (Arg17 and Arg76) are approximately 40 A apart, allowing the two reactive-site loops to bind to and to inhibit two trypsin molecules simultaneously and independently. The conformations of the reactive-site loops of BBBI are highly similar to those of other substrate-like inhibitors. This structure provides the framework for modeling of the 1:2 complex between BBBI and trypsin.
Organizational Affiliation:
College of Natural Sciences, Seoul National University, Seoul, 151-742, Korea.