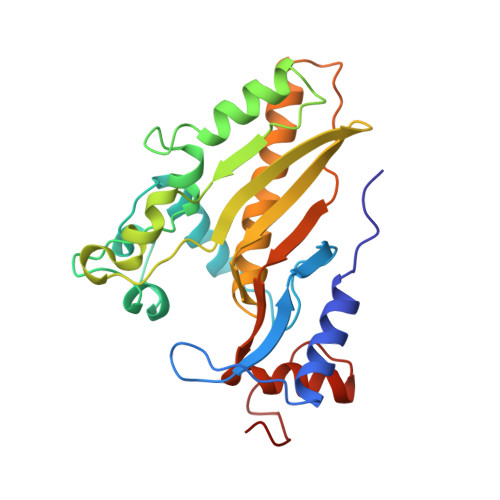
Crystal structure of deoxycytidylate hydroxymethylase from bacteriophage T4, a component of the deoxyribonucleoside triphosphate-synthesizing complex.
Song, H.K., Sohn, S.H., Suh, S.W.(1999) EMBO J 18: 1104-1113
- PubMed: 10064578
- DOI: https://doi.org/10.1093/emboj/18.5.1104
- Primary Citation of Related Structures:
1B49, 1B5D, 1B5E - PubMed Abstract:
Bacteriophage T4 deoxycytidylate hydroxymethylase (EC 2.1.2.8), a homodimer of 246-residue subunits, catalyzes hydroxymethylation of the cytosine base in deoxycytidylate (dCMP) to produce 5-hydroxymethyl-dCMP. It forms part of a phage DNA protection system and appears to function in vivo as a component of a multienzyme complex called deoxyribonucleoside triphosphate (dNTP) synthetase. We have determined its crystal structure in the presence of the substrate dCMP at 1.6 A resolution. The structure reveals a subunit fold and a dimerization pattern in common with thymidylate synthases, despite low (approximately 20%) sequence identity. Among the residues that form the dCMP binding site, those interacting with the sugar and phosphate are arranged in a configuration similar to the deoxyuridylate binding site of thymidylate synthases. However, the residues interacting directly or indirectly with the cytosine base show a more divergent structure and the presumed folate cofactor binding site is more open. Our structure reveals a water molecule properly positioned near C-6 of cytosine to add to the C-7 methylene intermediate during the last step of hydroxymethylation. On the basis of sequence comparison and crystal packing analysis, a hypothetical model for the interaction between T4 deoxycytidylate hydroxymethylase and T4 thymidylate synthase in the dNTP-synthesizing complex has been built.
Organizational Affiliation:
Department of Chemistry, College of Natural Sciences, Seoul National University, Seoul 151-742, Korea.