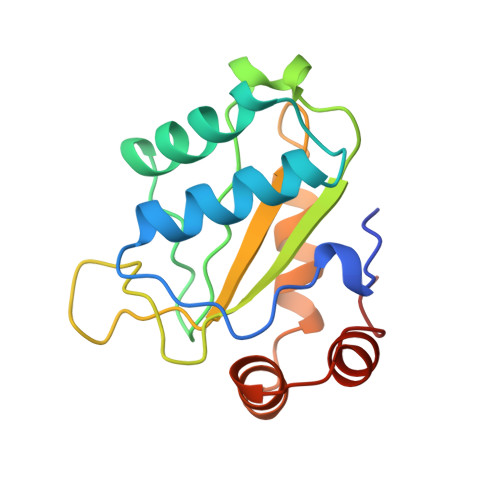
The structure of the Escherichia coli phosphotransferase IIAmannitol reveals a novel fold with two conformations of the active site.
van Montfort, R.L., Pijning, T., Kalk, K.H., Hangyi, I., Kouwijzer, M.L., Robillard, G.T., Dijkstra, B.W.(1998) Structure 6: 377-388
- PubMed: 9551558
- DOI: https://doi.org/10.1016/s0969-2126(98)00039-2
- Primary Citation of Related Structures:
1A3A - PubMed Abstract:
The bacterial phosphoenolpyruvate-dependent phosphotransferase system (PTS) catalyses the cellular uptake and subsequent phosphorylation of carbohydrates. Moreover, the PTS plays a crucial role in the global regulation of various metabolic pathways. The PTS consists of two general proteins, enzyme I and the histidine-containing protein (HPr), and the carbohydrate-specific enzyme II (EII). EIIs are usually composed of two cytoplasmic domains, IIA and IIB, and a transmembrane domain, IIC. The IIA domains catalyse the transfer of a phosphoryl group from HPr to IIB, which phosphorylates the transported carbohydrate. Knowledge of the structures of the IIA proteins may provide insight into the mechanisms by which the PTS couples phosphorylation reactions with carbohydrate specificity.
Organizational Affiliation:
Laboratory of Biophysical Chemistry, BIOSON Research Institute, University of Groningen, Nijenborgh, The Netherlands.