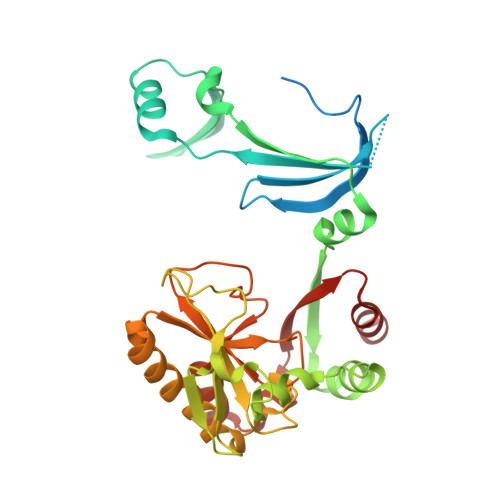
Crystal structures of human DcpS in ligand-free and m7GDP-bound forms suggest a dynamic mechanism for scavenger mRNA decapping.
Chen, N., Walsh, M.A., Liu, Y., Parker, R., Song, H.(2005) J Mol Biol 347: 707-718
- PubMed: 15769464
- DOI: https://doi.org/10.1016/j.jmb.2005.01.062
- Primary Citation of Related Structures:
1XML, 1XMM - PubMed Abstract:
Eukaryotic cells utilize DcpS, a scavenger decapping enzyme, to degrade the residual cap structure following 3'-5' mRNA decay, thereby preventing the premature decapping of the capped long mRNA and misincorporation of methylated nucleotides in nucleic acids. We report the structures of DcpS in ligand-free form and in a complex with m7GDP. apo-DcpS is a symmetric dimer, strikingly different from the asymmetric dimer observed in the structures of DcpS with bound cap analogues. In contrast, and similar to the m7GpppG-DcpS complex, DcpS with bound m7GDP is an asymmetric dimer in which the closed state appears to be the substrate-bound complex, whereas the open state mimics the product-bound complex. Comparisons of these structures revealed conformational changes of both the N-terminal swapped-dimeric domain and the cap-binding pocket upon cap binding. Moreover, Tyr273 in the cap-binding pocket displays remarkable conformational changes upon cap binding. Mutagenesis and biochemical analysis suggest that Tyr273 seems to play an important role in cap binding and product release. Examination of the crystallographic B-factors indicates that the N-terminal domain in apo-DcpS is inherently flexible, and in a dynamic state ready for substrate binding and product release.
Organizational Affiliation:
Laboratory of Macromolecular Structure, Institute of Molecular and Cell Biology, 61 Biopolis Drive, Proteos, Singapore 138673.