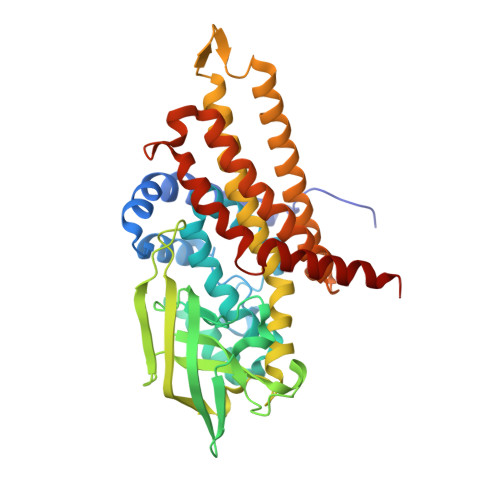
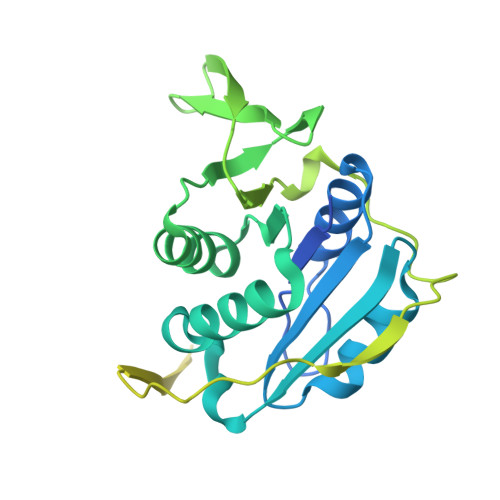
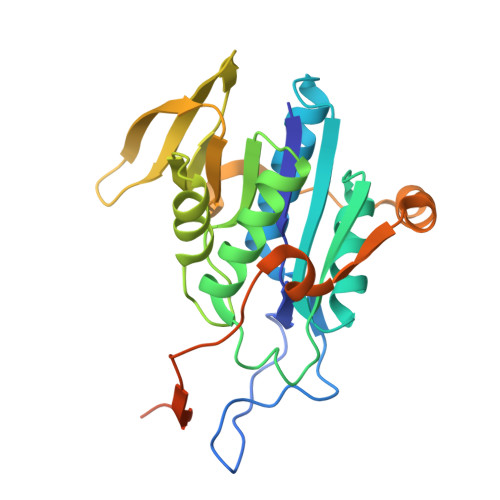
Extensive domain motion and electron transfer in the human electron transferring flavoprotein-medium chain Acyl-CoA dehydrogenase complex
Toogood, H.S., van Thiel, A., Basran, J., Sutcliffe, M.J., Scrutton, N.S., Leys, D.(2004) J Biol Chem 279: 32904-32912
- PubMed: 15159392
- DOI: https://doi.org/10.1074/jbc.M404884200
- Primary Citation of Related Structures:
1T9G - PubMed Abstract:
The crystal structure of the human electron transferring flavoprotein (ETF).medium chain acyl-CoA dehydrogenase (MCAD) complex reveals a dual mode of protein-protein interaction, imparting both specificity and promiscuity in the interaction of ETF with a range of structurally distinct primary dehydrogenases. ETF partitions the functions of partner binding and electron transfer between (i) the recognition loop, which acts as a static anchor at the ETF.MCAD interface, and (ii) the highly mobile redox active FAD domain. Together, these enable the FAD domain of ETF to sample a range of conformations, some compatible with fast interprotein electron transfer. Disorders in amino acid or fatty acid catabolism can be attributed to mutations at the protein-protein interface. Crucially, complex formation triggers mobility of the FAD domain, an induced disorder that contrasts with general models of protein-protein interaction by induced fit mechanisms. The subsequent interfacial motion in the MCAD.ETF complex is the basis for the interaction of ETF with structurally diverse protein partners. Solution studies using ETF and MCAD with mutations at the protein-protein interface support this dynamic model and indicate ionic interactions between MCAD Glu(212) and ETF Arg alpha(249) are likely to transiently stabilize productive conformations of the FAD domain leading to enhanced electron transfer rates between both partners.
Organizational Affiliation:
Department of Biochemistry, University of Leicester, Leicester LE1 7RH, United Kingdom.