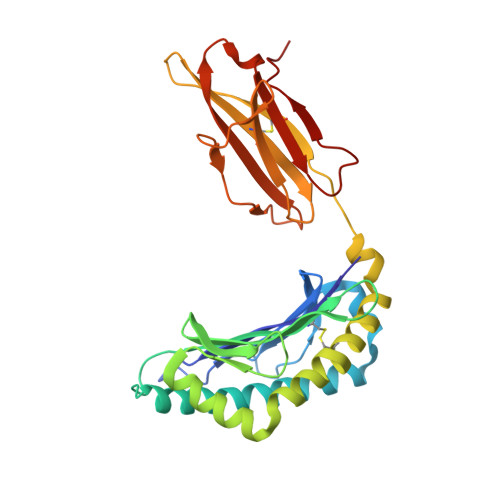
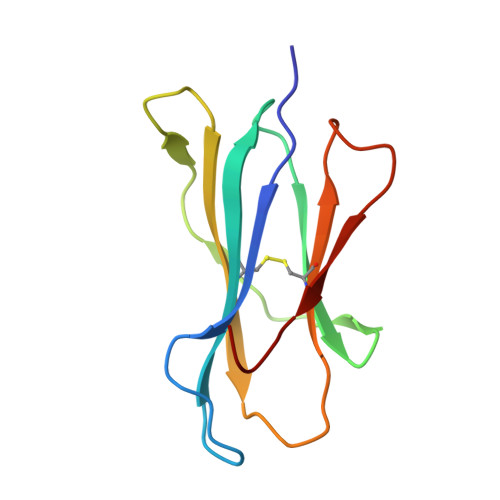
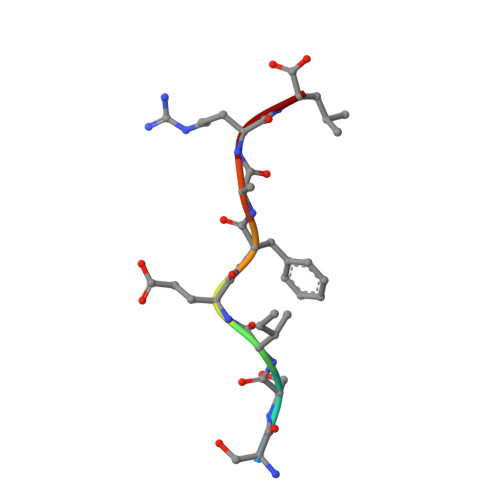
The structure of H-2K(b) and K(bm8) complexed to a herpes simplex virus determinant: evidence for a conformational switch that governs T cell repertoire selection and viral resistance.
Webb, A.I., Borg, N.A., Dunstone, M.A., Kjer-Nielsen, L., Beddoe, T., McCluskey, J., Carbone, F.R., Bottomley, S.P., Aguilar, M.I., Purcell, A.W., Rossjohn, J.(2004) J Immunol 173: 402-409
- PubMed: 15210799
- DOI: https://doi.org/10.4049/jimmunol.173.1.402
- Primary Citation of Related Structures:
1T0M, 1T0N - PubMed Abstract:
Polymorphism within the MHC not only affects peptide specificity but also has a critical influence on the T cell repertoire; for example, the CD8 T cell response toward an immunodominant HSV glycoprotein B peptide is more diverse and of higher avidity in H-2(bm8) compared with H-2(b) mice. We have examined the basis for the selection of these distinct antiviral T cell repertoires by comparing the high-resolution structures of K(b) and K(bm8), in complex with cognate peptide Ag. Although K(b) and K(bm8) differ by four residues within the Ag-binding cleft, the most striking difference in the two structures was the disparate conformation adopted by the shared residue, Arg(62). The altered dynamics of Arg(62), coupled with a small rigid-body movement in the alpha(1) helix encompassing this residue, correlated with biased Valpha usage in the B6 mice. Moreover, an analysis of all known TCR/MHC complexes reveals that Arg(62) invariably interacts with the TCR CDR1alpha loop. Accordingly, Arg(62) appears to function as a conformational switch that may govern T cell selection and protective immunity.
Organizational Affiliation:
Protein Crystallography Unit, Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Victoria 3800, Australia.