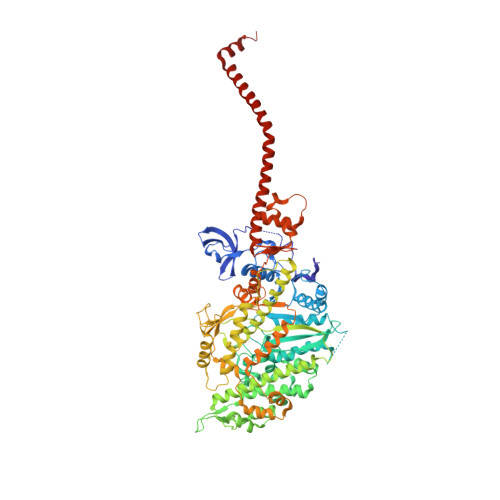
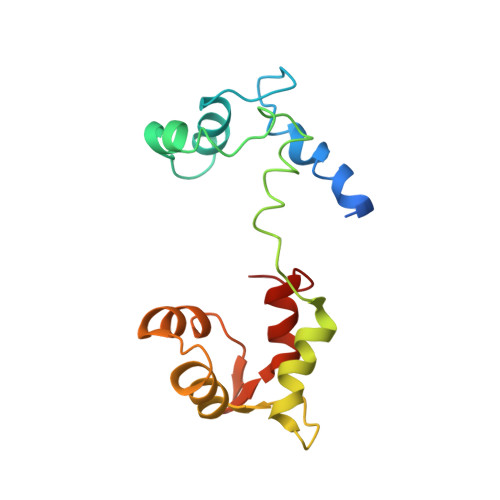
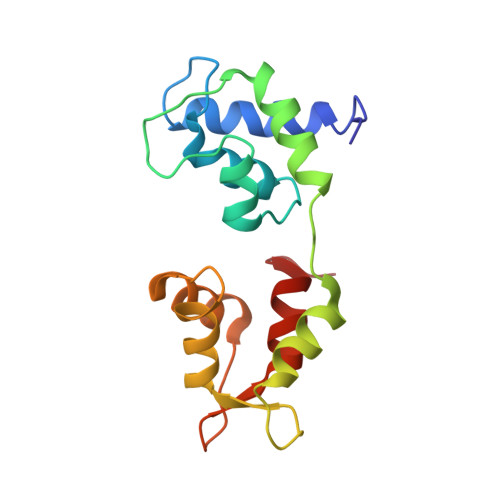
Myosin subfragment 1 structures reveal a partially bound nucleotide and a complex salt bridge that helps couple nucleotide and actin binding.
Risal, D., Gourinath, S., Himmel, D.M., Szent-Gyorgyi, A.G., Cohen, C.(2004) Proc Natl Acad Sci U S A 101: 8930-8935
- PubMed: 15184651
- DOI: https://doi.org/10.1073/pnas.0403002101
- Primary Citation of Related Structures:
1S5G, 1SR6 - PubMed Abstract:
Structural studies of myosin have indicated some of the conformational changes that occur in this protein during the contractile cycle, and we have now observed a conformational change in a bound nucleotide as well. The 3.1-A x-ray structure of the scallop myosin head domain (subfragment 1) in the ADP-bound near-rigor state (lever arm =45 degrees to the helical actin axis) shows the diphosphate moiety positioned on the surface of the nucleotide-binding pocket, rather than deep within it as had been observed previously. This conformation strongly suggests a specific mode of entry and exit of the nucleotide from the nucleotide-binding pocket through the so-called "front door." In addition, using a variety of scallop structures, including a relatively high-resolution 2.75-A nucleotide-free near-rigor structure, we have identified a conserved complex salt bridge connecting the 50-kDa upper and N-terminal subdomains. This salt bridge is present only in crystal structures of muscle myosin isoforms that exhibit a strong reciprocal relationship (also known as coupling) between actin and nucleotide affinity.
Organizational Affiliation:
Rosenstiel Basic Medical Sciences Research Center, MS 029, Waltham, MA 02454-9110, USA.