Cyanide binding to truncated hemoglobins: a crystallographic and kinetic study
Milani, M., Ouellet, Y., Ouellet, H., Guertin, M., Boffi, A., Antonini, G., Bocedi, A., Mattu, M., Bolognesi, M., Ascenzi, P.(2004) Biochemistry 43: 5213-5221
- PubMed: 15122887
- DOI: https://doi.org/10.1021/bi049870+
- Primary Citation of Related Structures:
1RTE - PubMed Abstract:
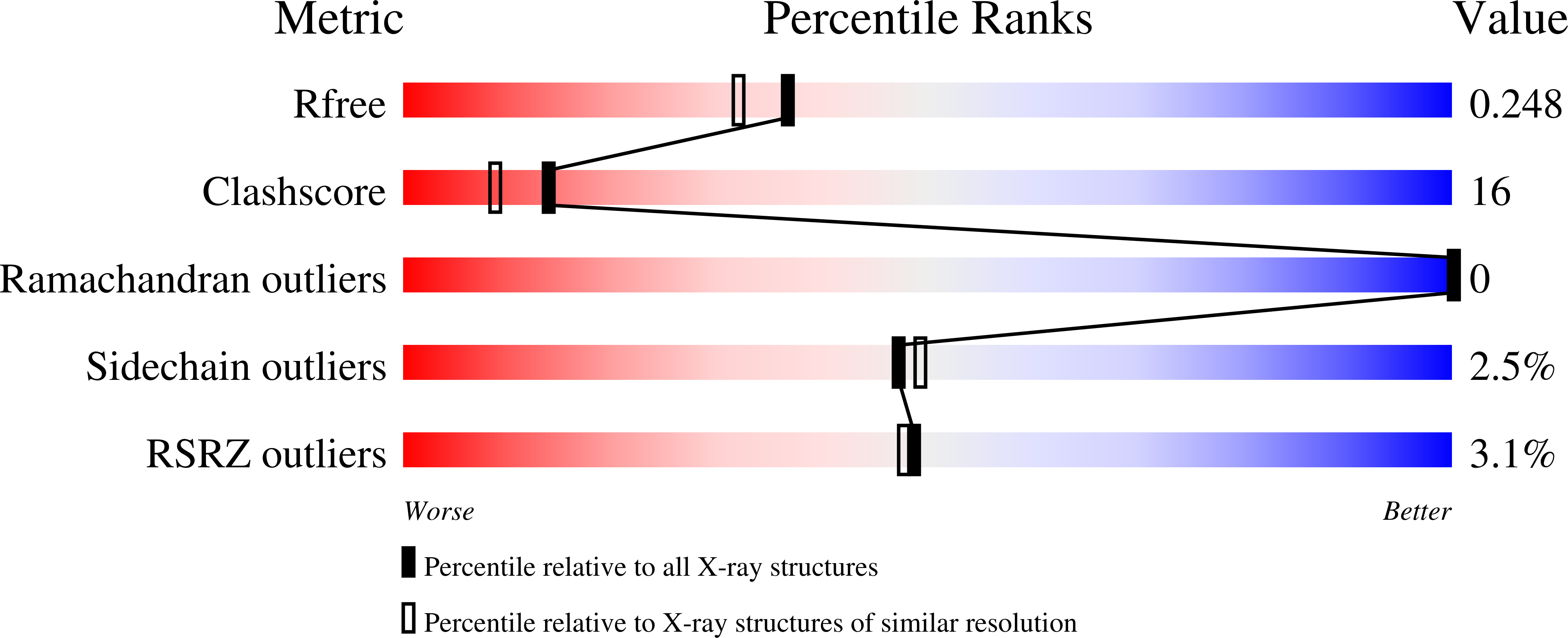
Cyanide is one of the few diatomic ligands able to interact with the ferric and ferrous heme-Fe atom. Here, the X-ray crystal structure of the cyanide derivative of ferric Mycobacterium tuberculosis truncated hemoglobin-N (M. tuberculosis trHbN) has been determined at 2.0 A (R-general = 17.8% and R-free = 23.5%), and analyzed in parallel with those of M. tuberculosis truncated hemoglobin-O (M. tuberculosis trHbO), Chlamydomonas eugametos truncated hemoglobin (C. eugametos trHb), and sperm whale myoglobin, generally taken as a molecular model. Cyanide binding to M. tuberculosis trHbN is stabilized directly by residue TyrB10(33), which may assist the deprotonation of the incoming ligand and the protonation of the outcoming cyanide. In M. tuberculosis trHbO and in C. eugametos trHb the ligand is stabilized by the distal pocket residues TyrCD1(36) and TrpG8(88), and by the TyrB10(20) - GlnE7(41) - GlnE11(45) triad, respectively. Moreover, kinetics for cyanide binding to ferric M. tuberculosis trHbN and trHbO and C. eugametos trHb, for ligand dissociation from the ferrous trHbs, and for the reduction of the heme-Fe(III)-cyanide complex have been determined, at pH 7.0 and 20.0 degrees C. Despite the different heme distal site structures and ligand interactions, values of the rate constant for cyanide binding to ferric (non)vertebrate heme proteins are similar, being influenced mainly by the presence in the heme pocket of proton acceptor group(s), whose function is to assist the deprotonation of the incoming ligand (i.e., HCN). On the other hand, values of the rate constant for the reduction of the heme-Fe(III)-cyanide (non)vertebrate globins span over several orders of magnitude, reflecting the different ability of the heme proteins considered to give productive complex(es) with dithionite or its reducing species SO(2)(-). Furthermore, values of the rate constant for ligand dissociation from heme-Fe(II)-cyanide (non)vertebrate heme proteins are very different, reflecting the different nature and geometry of the heme distal residue(s) hydrogen-bonded to the heme-bound cyanide.
Organizational Affiliation:
Istituto Giannina Gaslini, Largo G. Gaslini, 5. 16147 Genova, Italy.