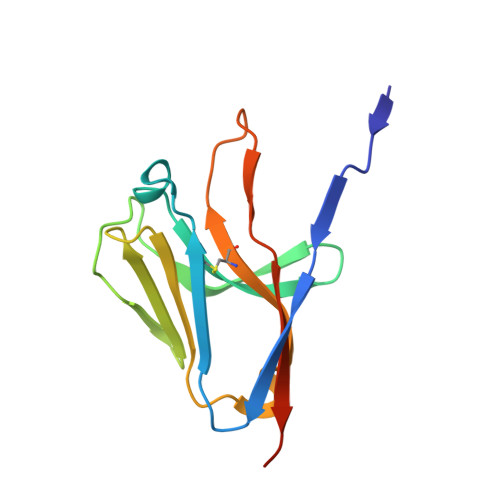
Crystal structure of the human myeloid cell activating receptor TREM-1
Radaev, S., Kattah, M., Rostro, B., Colonna, M., Sun, P.D.(2003) Structure 11: 1527-1535
- PubMed: 14656437
- DOI: https://doi.org/10.1016/j.str.2003.11.001
- Primary Citation of Related Structures:
1Q8M - PubMed Abstract:
Triggering receptors expressed on myeloid cells (TREM) are a family of recently discovered receptors that play important roles in innate immune responses, such as to activate inflammatory responses and to contribute to septic shock in response to microbial-mediated infections. To date, two TREM receptors in human and several homologs in mice have been identified. We report the 2.6 A resolution crystal structure of the extracellular domain of human TREM-1. The overall fold of the receptor resembles that of a V-type immunoglobulin domain with differences primarily located in the N-terminal strand. TREM-1 forms a "head-to-tail" dimer with 4100 A(2) interface area that is partially mediated by a domain swapping between the first strands. This mode of dimer formation is different from the "head-to-head" dimerization that existed in V(H)V(L) domains of antibodies or V domains of T cell receptors. As a result, the dimeric TREM-1 most likely contains two distinct ligand binding sites.
Organizational Affiliation:
Structural Immunology Section, Laboratory of Immunogenetics, National Institute of Allergy and Infectious Diseases, National Institutes of Health, 12441 Parklawn Drive, Rockville, MD 20852, USA.