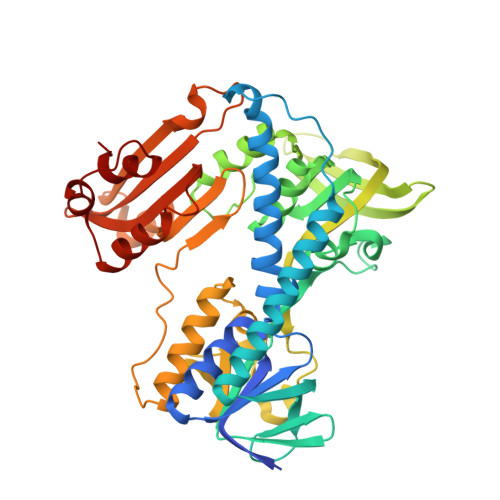
Substrate binding and catalysis by glutathione reductase as derived from refined enzyme: substrate crystal structures at 2 A resolution.
Karplus, P.A., Schulz, G.E.(1989) J Mol Biol 210: 163-180
- PubMed: 2585516
- DOI: https://doi.org/10.1016/0022-2836(89)90298-2
- Primary Citation of Related Structures:
1GRA, 1GRB, 1GRE, 1GRF, 1GRG - PubMed Abstract:
The X-ray structure analyses of four glutathione reductase complexes and derivatives have been extended to 2 A resolution and refined. The results are discussed in conjunction with the structure of the oxidized native enzyme known at 1.54 A resolution. While the residual co-ordinate errors are around 0.2 A, some significant shifts even in this range could be established. Points of particular interest are the 3.2 A approach of C4N of nicotinamide to N5F of flavin in hydride transfer geometry, the hydrogen bond geometries of the 2'-phosphate of NADPH as compared to inferior geometries for an inorganic phosphate binding together with NADH, the differential mobilities of parts of the substrates as derived from refined atomic temperature factors, and the stabilization of the thiolate of the proximal Cys63 by conformational changes of neighboring residues as well as by flavin. In addition, catalytically competent His467' is seen to interact more optimally with the sulfur of glutathione-I than with the distal sulfur of Cys58. The observed participation of water molecules for both NADPH and glutathione binding is so extensive that a prediction of the binding mode merely from the polypeptide structure would be very difficult. The accurately known geometries allowed us to draw some conclusions on the enzyme mechanism and suggest a possible scenario of the catalysis.
Organizational Affiliation:
Institut für Organische Chemie und Biochemie, Universität, Freiburg, F.R.G.