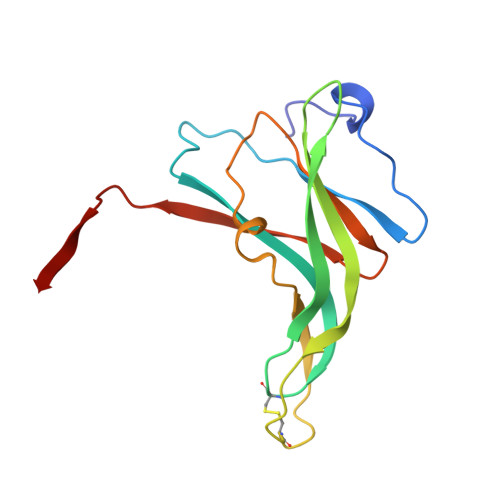
Crystal structure analyses of uncomplexed ecotin in two crystal forms: implications for its function and stability.
Shin, D.H., Song, H.K., Seong, I.S., Lee, C.S., Chung, C.H., Suh, S.W.(1996) Protein Sci 5: 2236-2247
- PubMed: 8931142
- DOI: https://doi.org/10.1002/pro.5560051110
- Primary Citation of Related Structures:
1ECY, 1ECZ - PubMed Abstract:
Ecotin, a homodimeric protein composed of 142 residue subunits, is a novel serine protease inhibitor present in Escherichia coli. Its thermostability and acid stability, as well as broad specificity toward proteases, make it an interesting protein for structural characterization. Its structure in the uncomplexed state, determined for two different crystalline environments, allows a structural comparison of the free inhibitor with that in complex with trypsin. Although there is no gross structural rearrangement of ecotin when binding trypsin, the loops involved in binding trypsin show relatively large shifts in atomic positions. The inherent flexibility of the loops and the highly nonglobular shape are the two features essential for its inhibitory function. An insight into the understanding of the structural basis of thermostability and acid stability of ecotin is also provided by the present structure.
Organizational Affiliation:
Department of Chemistry, Seoul National University, Korea.