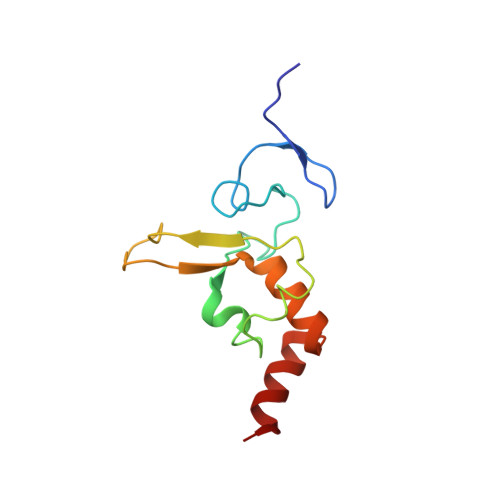
Interactions of human nucleotide excision repair protein XPA with DNA and RPA70 Delta C327: chemical shift mapping and 15N NMR relaxation studies.
Buchko, G.W., Daughdrill, G.W., de Lorimier, R., Rao B, K., Isern, N.G., Lingbeck, J.M., Taylor, J.S., Wold, M.S., Gochin, M., Spicer, L.D., Lowry, D.F., Kennedy, M.A.(1999) Biochemistry 38: 15116-15128
- PubMed: 10563794
- DOI: https://doi.org/10.1021/bi991755p
- Primary Citation of Related Structures:
1D4U - PubMed Abstract:
Human XPA is an essential component in the multienzyme nucleotide excision repair (NER) pathway. The solution structure of the minimal DNA binding domain of XPA (XPA-MBD: M98-F219) was recently determined [Buchko et al. (1998) Nucleic Acids Res. 26, 2779-2788, Ikegami et al. (1998) Nat. Struct. Biol. 5, 701-706] and shown to consist of a compact zinc-binding core and a loop-rich C-terminal subdomain connected by a linker sequence. Here, the solution structure of XPA-MBD was further refined using an entirely new class of restraints based on pseudocontact shifts measured in cobalt-substituted XPA-MBD. Using this structure, the surface of XPA-MBD which interacts with DNA and a fragment of the largest subunit of replication protein A (RPA70 Delta C327: M1-Y326) was determined using chemical shift mapping. DNA binding in XPA-MBD was highly localized in the loop-rich subdomain for DNA with or without a lesion [dihydrothymidine (dhT) or 6-4-thymidine-cytidine (64TC)], or with DNA in single- or double-stranded form, indicating that the character of the lesion itself is not the driving force for XPA binding DNA. RPA70 Delta C327 was found to contact regions in both the zinc-binding and loop-rich subdomains. Some overlap of the DNA and RPA70 Delta C327 binding regions was observed in the loop-rich subdomain, indicating a possible cooperative DNA-binding mode between XPA and RPA70 Delta C327. To complement the chemical shift mapping data, the backbone dynamics of free XPA-MBD and XPA-MBD bound to DNA oligomers containing dhT or 64TC lesions were investigated using 15N NMR relaxation data. The dynamic analyses for the XPA-MBD complexes with DNA revealed localized increases and decreases in S2 and an increase in the global correlation time. Regions of XPA-MBD with the largest increases in S2 overlapped regions having the largest chemical shifts changes upon binding DNA, indicating that the loop-rich subdomain becomes more rigid upon binding DNA. Interestingly, S2 decreased for some residues in the zinc-binding core upon DNA association, indicating a possible concerted structural rearrangement on binding DNA.
Organizational Affiliation:
Pacific Northwest National Laboratories, Environmental Molecular Sciences Laboratory, Richland, Washington 99352, USA.