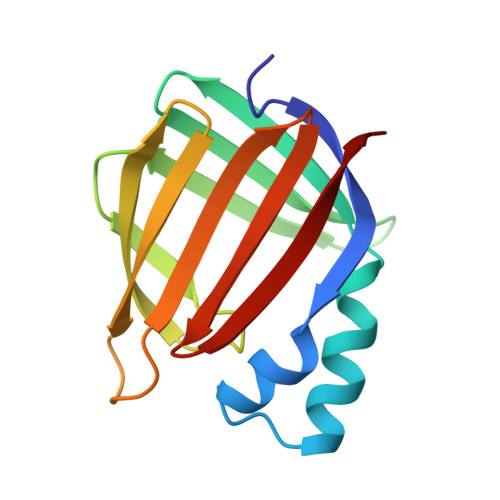
Crystal structure of cellular retinoic acid binding protein I shows increased access to the binding cavity due to formation of an intermolecular beta-sheet.
Thompson, J.R., Bratt, J.M., Banaszak, L.J.(1995) J Mol Biol 252: 433-446
- PubMed: 7563063
- DOI: https://doi.org/10.1006/jmbi.1995.0509
- Primary Citation of Related Structures:
1CBI - PubMed Abstract:
A recombinant form of murine apo-cellular retinoic acid binding protein I (apo-CRABPI) has been purified and crystallized at pH 5.0, and the crystal structure has been refined to an R-factor of 19.6% at a resolution of 2.7 A. CRABPI binds all-trans retinoic acid and some retinoic acid metabolites with nanomolar affinities. Coordinates of the holo form of CRABP were not available during the early stages of the study, and in spite of numerous homologs of known structure, phases were not obtainable through molecular replacement. Instead, an interpretable electron density map was obtained by multiple isomorphous replacement methods after improvement of the heavy-atom parameters with density modified trial phases. Two molecules of apo-CRABPI occupy the P3121 asymmetric unit and are related by pseudo 2-fold rotational symmetry. Unique conformational differences are apparent between the two molecules. In all of the family members studied to date, there is a lack of hydrogen bonds between two of the component beta-strands resulting in a gap in the interstand hydrogen bonding pattern. In the crystallographic dimer described here, a continuous intermolecular beta-sheet is formed by using this gap region. This is possible because of an 8 A outward maximum displacement of the tight turn between the third and fourth beta-strands on one of the molecules. The result is a double beta-barrel containing two apo-CRABPI molecules with a more open, ligand-accessible binding cavity, which has not been observed in other structures of a family of proteins that bind hydrophobic ligands.
Organizational Affiliation:
Department of Biochemistry, School of Medicine, University of Minnesota, Minneapolis 55455, USA.