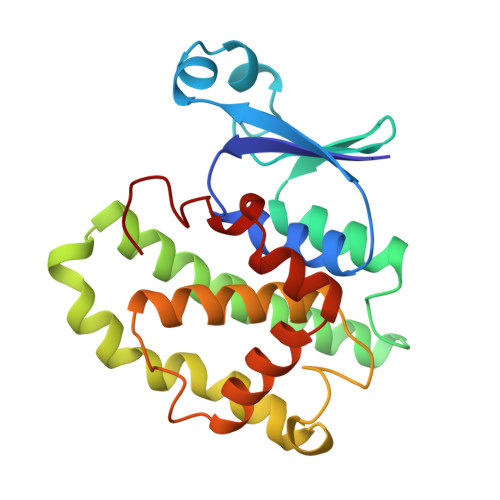
The glutathione conjugate of ethacrynic acid can bind to human pi class glutathione transferase P1-1 in two different modes.
Oakley, A.J., Lo Bello, M., Mazzetti, A.P., Federici, G., Parker, M.W.(1997) FEBS Lett 419: 32-36
- PubMed: 9426214
- DOI: https://doi.org/10.1016/s0014-5793(97)01424-5
- Primary Citation of Related Structures:
11GS - PubMed Abstract:
The diuretic drug ethacrynic acid, an inhibitor of pi class glutathione S-transferase, has been tested in clinical trials as an adjuvant in chemotherapy. We recently solved the crystal structure of this enzyme in complex with ethacrynic acid and its glutathione conjugate. Here we present a new structure of the ethacrynic-glutathione conjugate complex. In this structure the ethacrynic moiety of the complex is shown to bind in a completely different orientation to that previously observed. Thus there are at least two binding modes possible, an observation of great importance to the design of second generation inhibitors of the enzyme.
Organizational Affiliation:
The Ian Potter Foundation Protein Crystallography Laboratory, St. Vincent's Institute of Medical Research, Fitzroy, Vic., Australia.