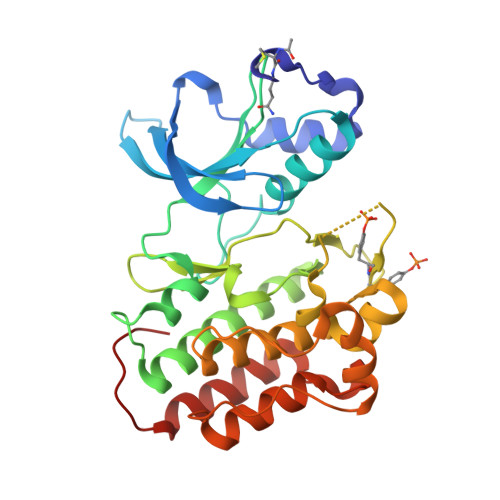
Discovery of 1-[3-(1-methyl-1H-pyrazol-4-yl)-5-oxo-5H-benzo[4,5]cyclohepta[1,2-b]pyridin-7-yl]-N-(pyridin-2-ylmethyl)methanesulfonamide (MK-8033): A Specific c-Met/Ron dual kinase inhibitor with preferential affinity for the activated state of c-Met.
Northrup, A.B., Katcher, M.H., Altman, M.D., Chenard, M., Daniels, M.H., Deshmukh, S.V., Falcone, D., Guerin, D.J., Hatch, H., Li, C., Lu, W., Lutterbach, B., Allison, T.J., Patel, S.B., Reilly, J.F., Reutershan, M., Rickert, K.W., Rosenstein, C., Soisson, S.M., Szewczak, A.A., Walker, D., Wilson, K., Young, J.R., Pan, B.S., Dinsmore, C.J.(2013) J Med Chem 56: 2294-2310
- PubMed: 23379595
- DOI: https://doi.org/10.1021/jm301619u
- Primary Citation of Related Structures:
4IWD - PubMed Abstract:
This report documents the first example of a specific inhibitor of protein kinases with preferential binding to the activated kinase conformation: 5H-benzo[4,5]cyclohepta[1,2-b]pyridin-5-one 11r (MK-8033), a dual c-Met/Ron inhibitor under investigation as a treatment for cancer. The design of 11r was based on the desire to reduce time-dependent inhibition of CYP3A4 (TDI) by members of this structural class. A novel two-step protocol for the synthesis of benzylic sulfonamides was developed to access 11r and analogues. We provide a rationale for the observed selectivity based on X-ray crystallographic evidence and discuss selectivity trends with additional examples. Importantly, 11r provides full inhibition of tumor growth in a c-Met amplified (GTL-16) subcutaneous tumor xenograft model and may have an advantage over inactive form kinase inhibitors due to equal potency against a panel of oncogenic activating mutations of c-Met in contrast to c-Met inhibitors without preferential binding to the active kinase conformation.
Organizational Affiliation:
Department of Chemistry, Merck & Co., Inc. , 33 Avenue Louis Pasteur, BMB-3, Boston, Massachusetts 02115, USA. alan_northrup@merck.com