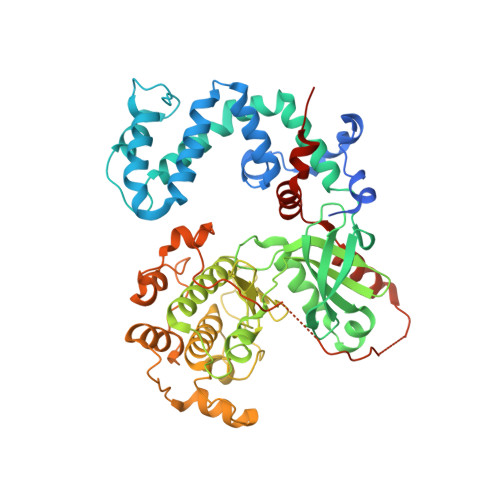
Structure of a monomeric variant of rhodopsin kinase at 2.5 A resolution.
Tesmer, J.J., Nance, M.R., Singh, P., Lee, H.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 622-625
- PubMed: 22684056
- DOI: https://doi.org/10.1107/S1744309112017435
- Primary Citation of Related Structures:
3T8O - PubMed Abstract:
G protein-coupled receptor kinase 1 (GRK1 or rhodopsin kinase) phosphorylates activated rhodopsin and initiates a cascade of events that results in the termination of phototransduction by the receptor. Although GRK1 seems to be a monomer in solution, seven prior crystal structures of GRK1 revealed a similar domain-swapped dimer interface involving the C-terminus of the enzyme. The influence of this interface on the overall conformation of GRK1 is not known. To address this question, the crystalline dimer interface was disrupted with a L166K mutation and the structure of GRK1-L166K was determined in complex with Mg(2+) · ATP to 2.5 Å resolution. GRK1-L166K crystallized in a novel space group as a monomer and exhibited little overall conformational difference from prior structures of GRK1, although the C-terminal domain-swapped region had reorganized owing to loss of the dimer interface.
Organizational Affiliation:
Life Sciences Institute and the Department of Pharmacology, University of Michigan, 210 Washtenaw Avenue, Ann Arbor, MI 48109-2216, USA. tesmerjj@umich.edu