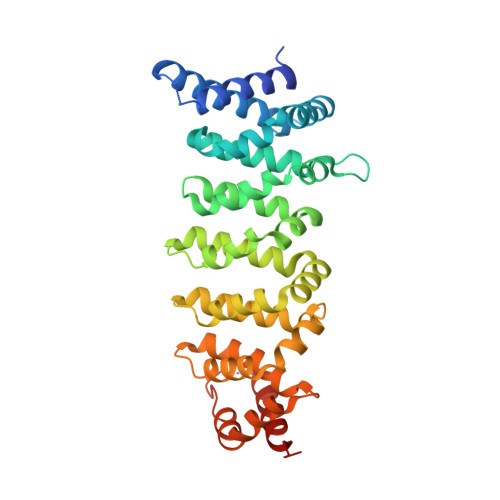
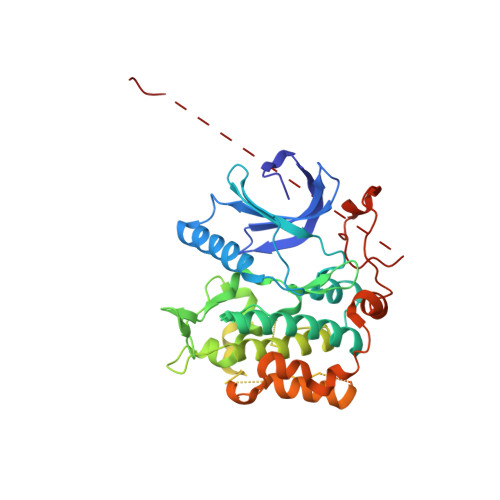
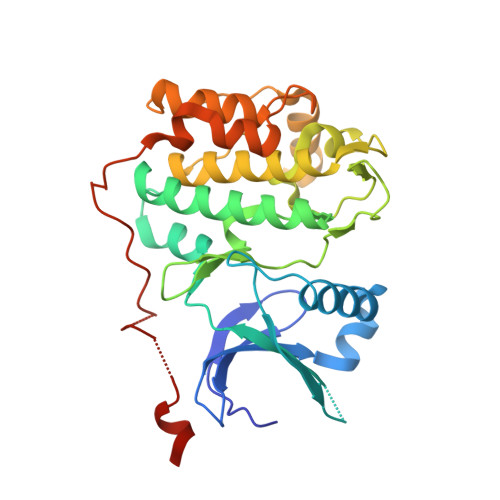
Structure of the Lkb1-Strad-Mo25 Complex Reveals an Allosteric Mechanism of Kinase Activation.
Zeqiraj, E., Filippi, B.M., Deak, M., Alessi, D.R., Van Aalten, D.M.F.(2009) Science 326: 1707
- PubMed: 19892943
- DOI: https://doi.org/10.1126/science.1178377
- Primary Citation of Related Structures:
2WTK - PubMed Abstract:
The LKB1 tumor suppressor is a protein kinase that controls the activity of adenosine monophosphate-activated protein kinase (AMPK). LKB1 activity is regulated by the pseudokinase STRADalpha and the scaffolding protein MO25alpha through an unknown, phosphorylation-independent, mechanism. We describe the structure of the core heterotrimeric LKB1-STRADalpha-MO25alpha complex, revealing an unusual allosteric mechanism of LKB1 activation. STRADalpha adopts a closed conformation typical of active protein kinases and binds LKB1 as a pseudosubstrate. STRADalpha and MO25alpha promote the active conformation of LKB1, which is stabilized by MO25alpha interacting with the LKB1 activation loop. This previously undescribed mechanism of kinase activation may be relevant to understanding the evolution of other pseudokinases. The structure also reveals how mutations found in Peutz-Jeghers syndrome and in various sporadic cancers impair LKB1 function.
Organizational Affiliation:
Division of Molecular Microbiology, College of Life Sciences, University of Dundee, Dundee DD1 5EH, Scotland.