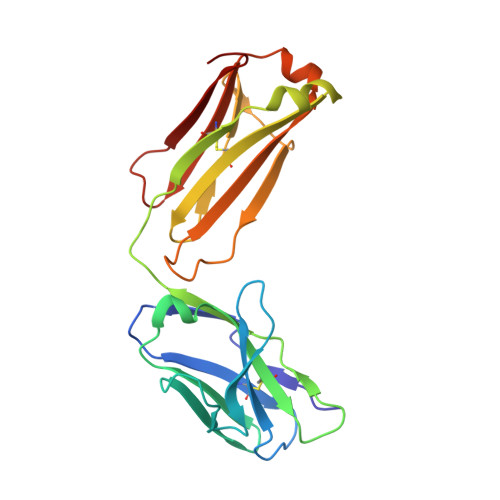
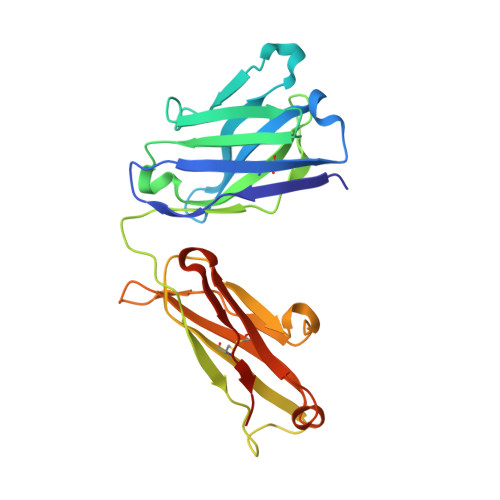
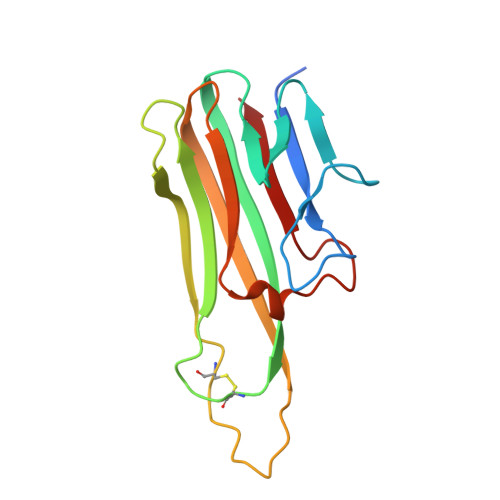
Structural basis for treating tumor necrosis factor alpha (TNFalpha)-associated diseases with the therapeutic antibody infliximab
Liang, S.Y., Dai, J.X., Hou, S., Su, L., Zhang, D., Guo, H., Hu, S., Wang, H., Rao, Z., Guo, Y.J., Lou, Z.Y.(2013) J Biol Chem 288: 13799-13807
- PubMed: 23504311
- DOI: https://doi.org/10.1074/jbc.M112.433961
- Primary Citation of Related Structures:
4G3Y - PubMed Abstract:
Although infliximab has high efficacy in treating TNFα-associated diseases, the epitope on TNFα remains unclear.
Organizational Affiliation:
Laboratory of Structural Biology and Ministry of Education Laboratory of Protein Science, School of Medicine, Tsinghua University, Beijing 100084, China.