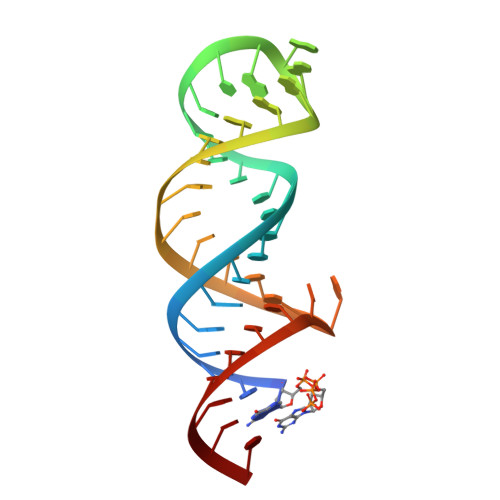
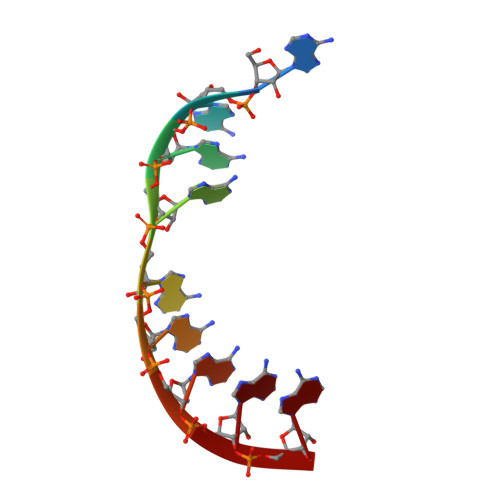
Poly(A) tail recognition by a viral RNA element through assembly of a triple helix.
Mitton-Fry, R.M., DeGregorio, S.J., Wang, J., Steitz, T.A., Steitz, J.A.(2010) Science 330: 1244-1247
- PubMed: 21109672
- DOI: https://doi.org/10.1126/science.1195858
- Primary Citation of Related Structures:
3P22 - PubMed Abstract:
Kaposi's sarcoma-associated herpesvirus produces a highly abundant, nuclear noncoding RNA, polyadenylated nuclear (PAN) RNA, which contains an element that prevents its decay. The 79-nucleotide expression and nuclear retention element (ENE) was proposed to adopt a secondary structure like that of a box H/ACA small nucleolar RNA (snoRNA), with a U-rich internal loop that hybridizes to and protects the PAN RNA poly(A) tail. The crystal structure of a complex between the 40-nucleotide ENE core and oligo(A)(9) RNA at 2.5 angstrom resolution reveals that unlike snoRNAs, the U-rich loop of the ENE engages its target through formation of a major-groove triple helix. A-minor interactions extend the binding interface. Deadenylation assays confirm the functional importance of the triple helix. Thus, the ENE acts as an intramolecular RNA clamp, sequestering the PAN poly(A) tail and preventing the initiation of RNA decay.
Organizational Affiliation:
Department of Molecular Biophysics and Biochemistry (MB&B), Howard Hughes Medical Institute (HHMI), Yale University School of Medicine, Boyer Center for Molecular Medicine, 295 Congress Avenue, New Haven, CT 06536-9812, USA.