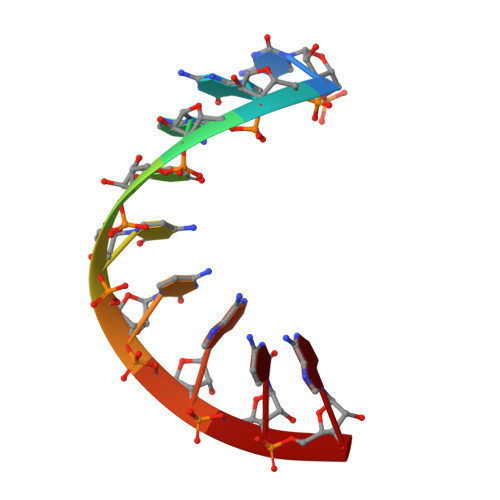
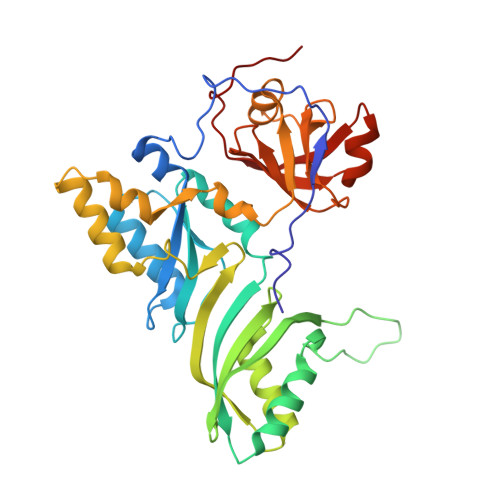
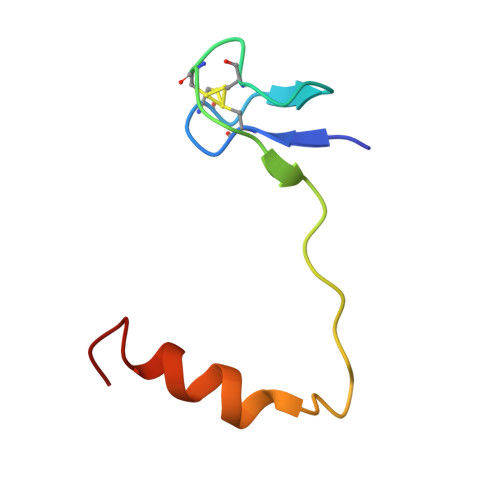
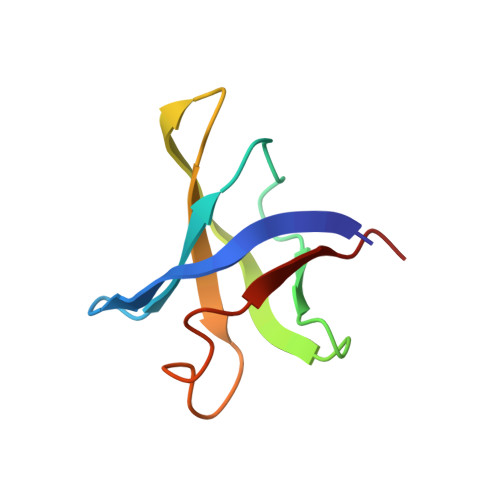
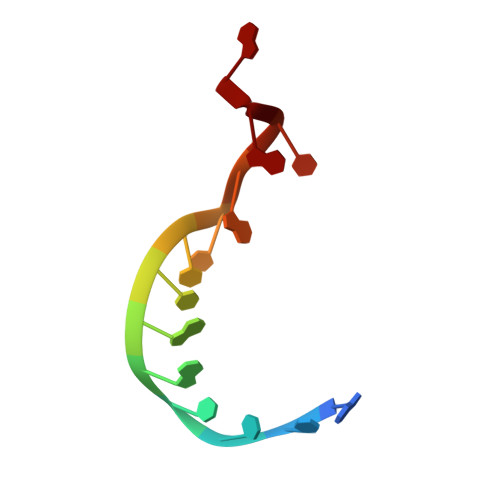
Structural and functional evidence of high specificity of Cbf5 for ACA trinucleotide.
Zhou, J., Liang, B., Li, H.(2011) RNA 17: 244-250
- PubMed: 21149572
- DOI: https://doi.org/10.1261/rna.2415811
- Primary Citation of Related Structures:
3MQK - PubMed Abstract:
Cbf5 is the catalytic subunit of the H/ACA small nucleolar/Cajal body ribonucleoprotein particles (RNPs) responsible for site specific isomerization of uridine in ribosomal and small nuclear RNA. Recent evidence from studies on archaeal Cbf5 suggests its second functional role in modifying tRNA U55 independent of guide RNA. In order to act both as a stand-alone and a RNP pseudouridine synthase, Cbf5 must differentiate features in H/ACA RNA from those in tRNA or rRNA. Most H/ACA RNAs contain a hallmark ACA trinucleotide downstream of the H/ACA motif. Here we challenged an archaeal Cbf5 (in the form of a ternary complex with its accessory proteins Nop10 and Gar1) with T-stem-loop RNAs with or without ACA trinucleotide in the stem. Although these substrates were previously shown to be substrates for the bacterial stand-alone pseudouridine synthase TruB, the Cbf5-Nop10-Gar1 complex was only able to modify those without ACA trinucleotide. A crystal structure of Cbf5-Nop10-Gar1 trimer bound with an ACA-containing T-stem-loop revealed that the ACA trinucleotide detracted Cbf5 from the stand-alone binding mode, thereby suggesting that the H/ACA RNP-associated function of Cbf5 likely supersedes its stand-alone function.
Organizational Affiliation:
Department of Chemistry and Biochemistry, Florida State University, Tallahassee, Florida 32306, USA.