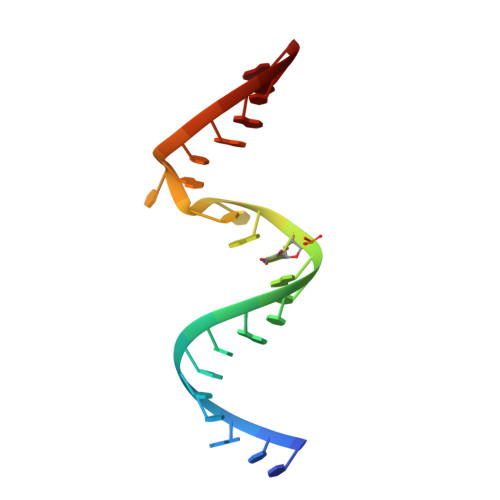
Structure of an RNA dimer of a regulatory element from human thymidylate synthase mRNA.
Dibrov, S., McLean, J., Hermann, T.(2011) Acta Crystallogr D Biol Crystallogr 67: 97-104
- PubMed: 21245530
- DOI: https://doi.org/10.1107/S0907444910050900
- Primary Citation of Related Structures:
3MEI - PubMed Abstract:
A sequence around the start codon of the mRNA of human thymidylate synthase (TS) folds into a secondary-structure motif in which the initiation site is sequestered in a metastable hairpin. Binding of the protein to its own mRNA at the hairpin prevents the production of TS through a translation-repression feedback mechanism. Stabilization of the mRNA hairpin by other ligands has been proposed as a strategy to reduce TS levels in anticancer therapy. Rapidly proliferating cells require high TS activity to maintain the production of thymidine as a building block for DNA synthesis. The crystal structure of a model oligonucleotide (TS1) that represents the TS-binding site of the mRNA has been determined. While fluorescence studies showed that the TS1 RNA preferentially adopts a hairpin structure in solution, even at high RNA concentrations, an asymmetric dimer of two hybridized TS1 strands was obtained in the crystal. The TS1 dimer contains an unusual S-turn motif that also occurs in the `off' state of the human ribosomal decoding site RNA.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of California, San Diego, 9500 Gilman Drive, La Jolla, CA 92093, USA.